-
-

The INSILICO Experience
-
Meet Whobee
-
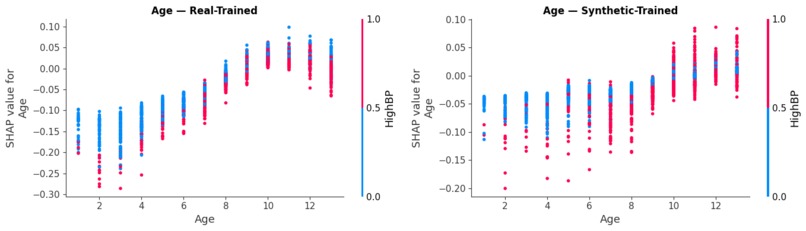
Biological Consistency: Side-by-side dependence plots showing how diabetes risk scales accurately with age in our digital twin cohort.
-
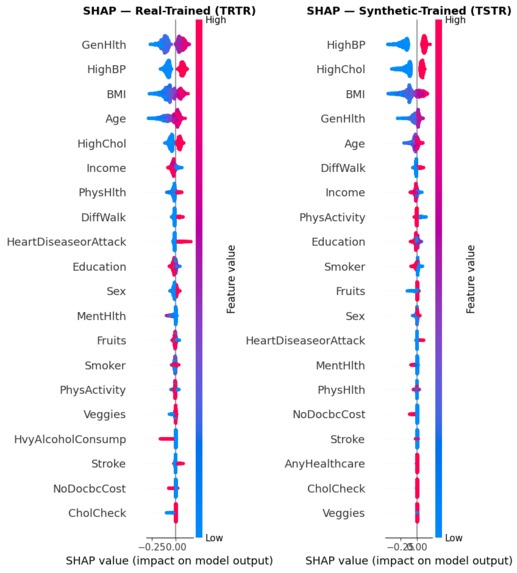
Comparing SHAP beeswarms to ensure our synthetic-trained model mirrors the clinical logic of real-world data.
-
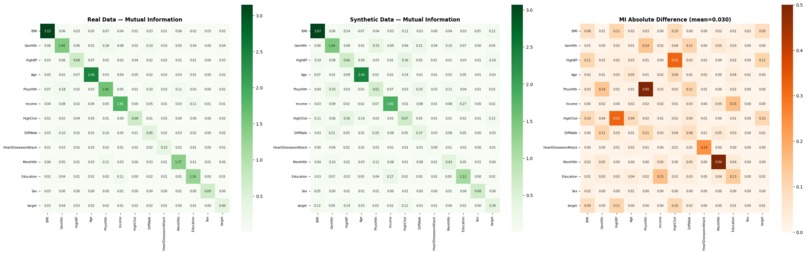
Mutual Information fidelity maps (mean diff: 0.029) confirm the "hidden DNA" of health markers is preserved.
-
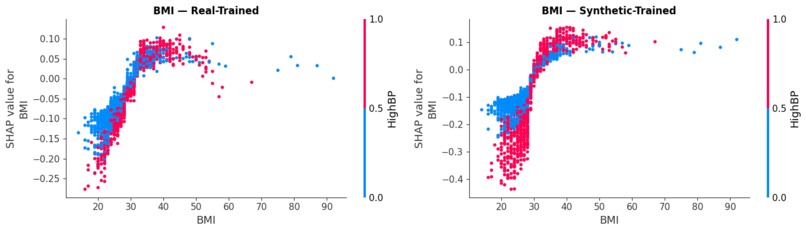
Dependence plots illustrating how BMI and HighBP interact to drive risk, validating our model's non-linear reasoning.
-
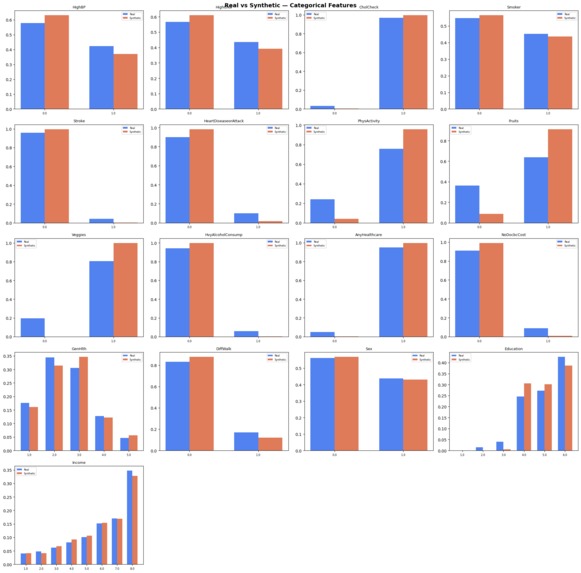
High-fidelity distribution matching across 10,000 patients, proving our TVAE successfully cloned the BRFSS cohort.
Inspiration
Nine out of ten drugs that enter a clinical trial never make it out. Each failure burns hundreds of millions of dollars and, more painfully, years of patient lives we don't get back. We kept asking the same question: why is the first time we learn a drug won't work always a real person?
Two things changed recently that made us think we could do something about it. LLMs got good enough to read a 50-page trial PDF and pull real numbers out of it. Synthetic data generators got good enough to make fake patients that still behave like real ones. Neither is useful alone. Glued together, they let you run a trial in silico before you run it on anyone.
What it does
You upload a clinical trial PDF. InSilico pulls the drug's expected effects out of it, distills them into a single strength score we call it gamma, between 0 and 1 and runs the drug against 720 simulated patient twins built around the profile you enter. Takes about 30 seconds.
The output is a cohort response: how many of those 720 patients responded, partially responded, didn't respond, or got worse plus per-patient SHAP explainability for the risk shift. Analysts get a go / no-go signal before they spend real money.
Alongside the simulator, we built Whobee, a patient-facing companion chatbot. The simulator is for the analyst deciding whether a trial is worth running. Whobee is for the patient actually enrolled in it "what do I do if I miss my dose?", "explain my HbA1c number" answered in plain language, grounded in the trial's own protocol.
How we built it
The pipeline has six moving parts:
Ingest & RAG. The PDF is chunked, embedded with Gemini, and stored in a per-request ChromaDB collection. We run six targeted queries HbA1c, weight, blood pressure, relative risk reduction, trial metadata, mechanism and pull the relevant chunks.
Endpoint extraction. Gemini reads those chunks and returns the actual measured numbers
Gamma formula. If at least two of the four key endpoints came out cleanly, we plug them into a calibrated linear form where the weights were hand-picked from ten reference drugs (semaglutide, tirzepatide, metformin, and peers). If fewer than two endpoints show up, we fall back to asking Gemini for gamma directly and label it llm_guess in the UI.
Synthetic cohort. Gamma drives a TVAE trained on the CDC BRFSS 2015 dataset originally 253,000 patients, randomly sampled to 10,000 for tractable training. We draw 720 twins around the input patient profile.
Risk scoring & explainability. A RandomForest scores baseline and post-drug risk for every twin; SHAP runs on top for per-patient explanations. A Next.js 15 frontend renders the trajectory, cohort pie, local SHAP, and feature deltas.
Whobee. Built end-to-end with Open Claw. The hard part of a medical chatbot isn't making it talk it's making it safe. Open Claw drafted the guardrails, the refusal logic for anything that looks like dosing advice beyond protocol, and the session memory that remembers what the patient already said.
Challenges we ran into
Gemini rate-limits. We hit the free-tier ceiling mid-testing. We had to design the pipeline to degrade gracefully the llm_guess path exists because of this, and the UI had to be honest about when we were measuring versus guessing. Qualitative PDFs. Consent forms and protocol synopses use language like "rapid HbA1c reduction" with no numbers. The formula gate skips them by design but teaching the UI to label that clearly, instead of silently falling back, took more iteration than the math did. TVAE training time vs. realism. The full 253k-patient BRFSS dataset trained a better generator but made the pipeline too slow for a live demo. Sampling down to 10k preserved the correlations we cared about and kept end-to-end under 35 seconds. Coefficient calibration with no ground truth. We had no "correct" gamma to fit against, so the ten reference drugs had to be picked carefully trials whose outcomes are well-documented across populations, not just one study.
Accomplishments that we're proud of
End-to-end in 30 seconds. No mocked stages. The PDF actually gets read, the twins actually get generated, SHAP actually runs. Honest fallbacks. When the system can't measure something, it says so. llm_guess is visible to the user. That felt more important than hiding it. Whobee. A patient-safety-aware companion, built in hours with Open Claw, that we'd actually feel okay putting in front of a real trial participant. The label change. Halfway through the build we renamed the project from TrialForge to InSilico it said more clearly what the thing is.
What we learned
LLMs are great at extraction, bad at arithmetic. Letting Gemini compute gamma directly was tempting. It was also wrong often enough that the formula-first / LLM-second ordering mattered. Synthetic data preserves correlations, not individuals. Our twins aren't "this patient" they're "a patient like this." Communicating that in the UI was as important as the modeling. Small datasets, picked well, beat big datasets, picked lazily. 10k randomly-sampled BRFSS rows captured the structure we needed; adding more rows did not make the generator meaningfully better on a weekend timeline. Agents earn trust by being boring. Open Claw didn't write flashy code it wrote the guardrails, the fallbacks, the labelling. That's what made Whobee ship-able.
What's next for InSilico
Confidence intervals on gamma clinicians should see uncertainty, not false precision. Cohort size as a knob 720 is a nice demo number; phase-I asks and phase-III asks aren't the same question. Auto-fit coefficients the ten hand-picked weights become a learned regression over a growing reference library as new trial readouts land. Multi-arm trials, drug-drug interactions, genomic stratification so we can answer "what would this drug do to a BRCA-positive sub-cohort" instead of an average patient. Real-EHR integration + audit trails the path toward the FDA's Model-Informed Drug Development program, where simulated cohorts are becoming admissible evidence. Whobee, multilingual and schedule-aware so "when is my next visit?" stops being a demo answer and becomes a real one. The horizon is a world where every trial is simulated before it's funded, and every patient in one has someone to talk to at 2 a.m.
Built With
- chromadb
- fastapi
- gemini
- html5
- next.js
- numpy
- pandas
- python
- radixui
- rag
- scikit-learn
- sdv
- shap
- spline
- tailwind
- typescript







Log in or sign up for Devpost to join the conversation.