-
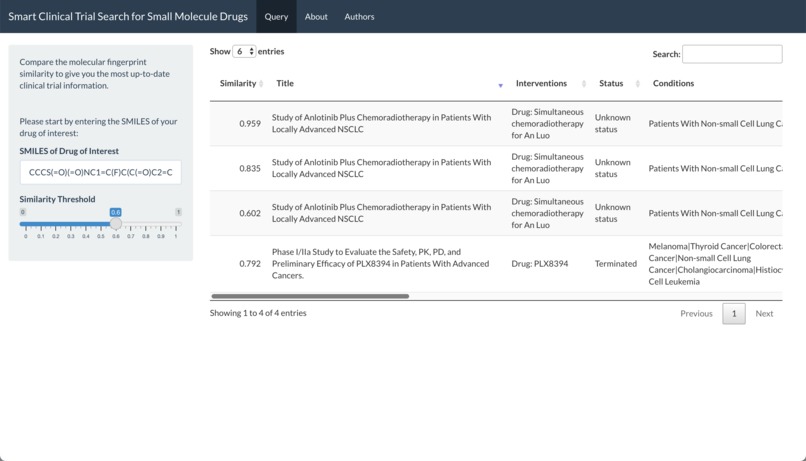
Querying with SMILES of Vemurafenib (1): 4 clinical trials on drugs with similarity > 0.6 are found
-

Querying with SMILES of Vemurafenib (2): 4 clinical trials on drugs with similarity > 0.6 are found
-

Querying with SMILES of Vemurafenib (3): 4 clinical trials on drugs with similarity > 0.6 are found
-

Querying with SMILES of Gefitinib, sorted by trial status to show completed clinical trials
-

The About page
-

The Authors page
-

Project architecture
Inspiration
- Just found out an existing small molecule drug with the potential to enter the clinical trial for ANOTHER disease?
- Just designed a BRAND NEW small molecule with the potential to better target a disease?
- It's difficult and tedious to search for complete clinical trial information across papers and databases without correct search keywords.
What researchers can get with our app
- A brand new drug with no name / An old drug but considering it for other disease targets?
- No problem, we will search for similar drugs in all clinical trials for various diseases for you.
- You can see what other organizations have done with your drug or similar drugs (Completed/Ongoing/Recruiting clinical trials and their results), and get a huge blueprint on the current drug discovery research.
How we built it
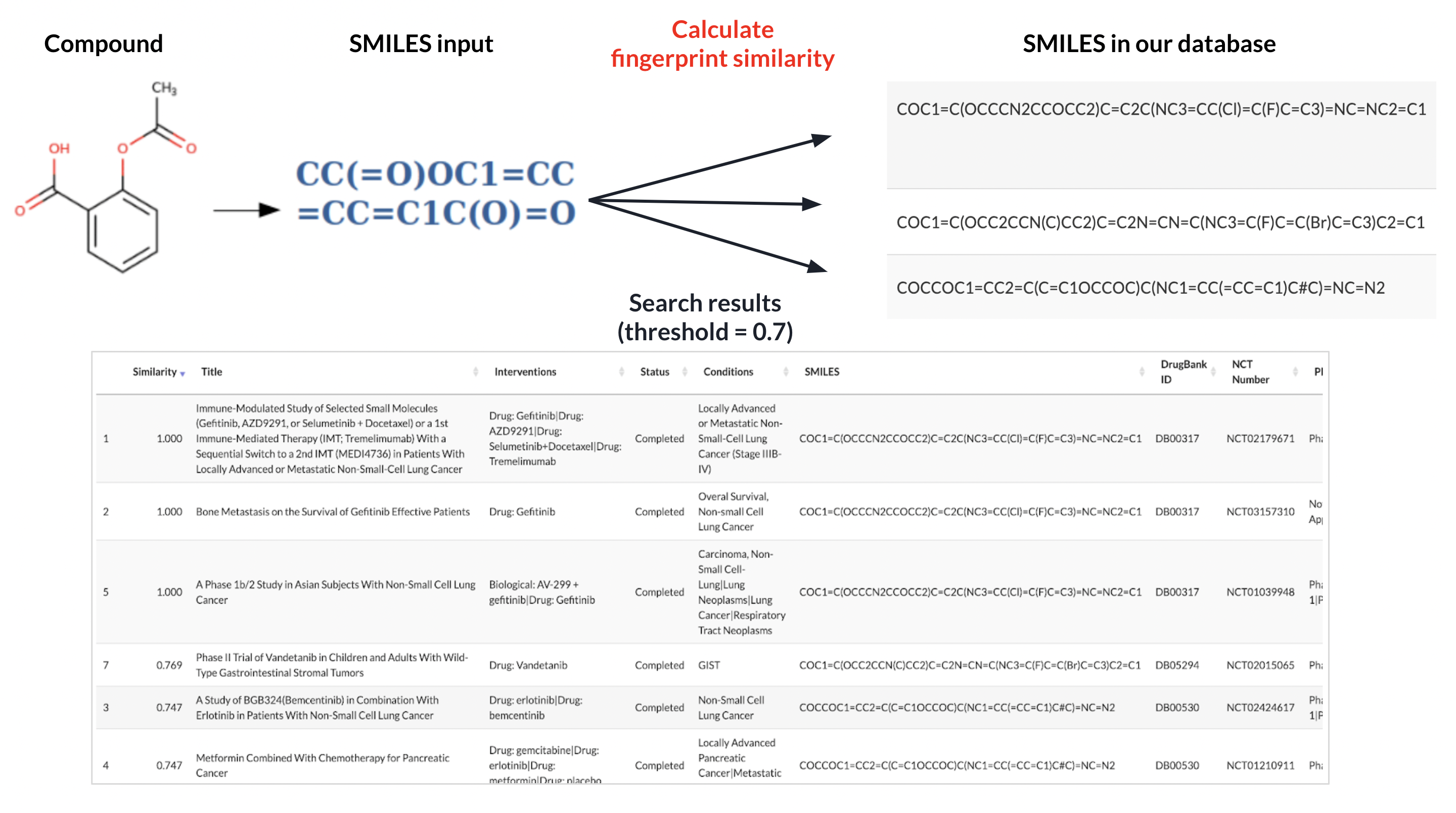
SMILES (simplified molecular-input line-entry system) is a specification in the form of a line notation for describing the structure of chemical species using short strings. Using SMILES, we can calculate the molecular structural similarity between different drugs. We search several largest clinical trial and drug databases including ClinicalTrials.gov and DrugBank, pre-process, clean, and merge those data to retrieve the information we need for calculating SMILES similarity. Then we use RDKit to calculate molecular fingerprint similarity between a query drug and all drugs in the database, and the information of a list of clinical trials on similar drugs can thus be obtained.
Challenges we ran into
The dataset pre-processing, cleaning, and merging steps, as well as the molecular similarity calculation step, are all done in Python, while the user interface of our app is implemented using Shiny in R. The interactive functionality of acquiring user inputs, calculating similarities, and querying the database require connecting Python packages and scripts to R, and we had some difficulties with this in the beginning.
Accomplishments that we're proud of
- We designed the very first search engine for clinical trials based on molecular fingerprint similarity calculation.
- No chemical name input is required from the user. Just compute the SMILES in any software and we will do the rest.
- We linked multiple databases through data wrangling and cleaning, leveraged R Shiny App to make the UI interface driven by the in-time user data input, and successfully integrated Python packages and scripts into the R programming environment.
What we learned
- RDKit calculation for molecular similarity (Python only)
- Integrating RDKit python script into the R environment
- Developing front-end and back-end algorithms in Shiny app
- Data wrangling and cleaning skills given the complex data across databases
What's next for Smart Clinical Trial Search for Small Molecule Drugs
More functionalities like:
- Merging more clinical trial databases (Ex. by Web Crawling)
- Adding more user control in the R shiny app UI (Ex. clinical trial status, search by drug name)
- Generating images of drug structures
- Generating more plots to demonstrate the molecule/drug relationships and interactions
Built With
- clinicaltrials.gov
- drugbank
- google-colab
- python
- r
- rdkit
- shiny







Log in or sign up for Devpost to join the conversation.