-
-
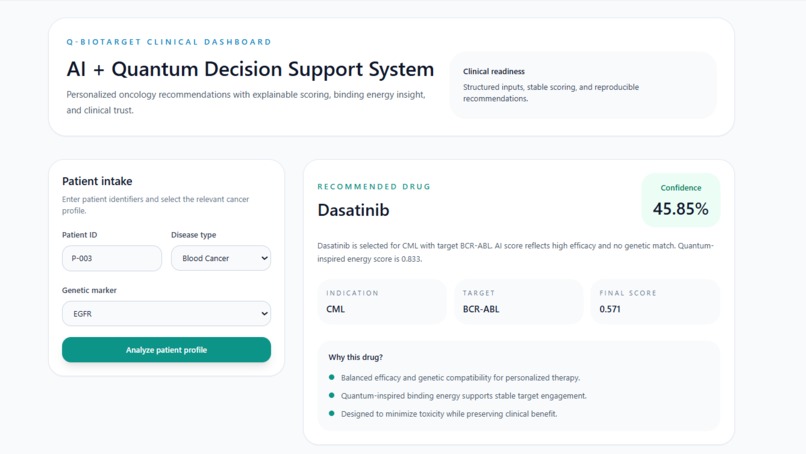
Enter Patient ID, Disease Type & Genetic Marker Q-BioTarget instantly recommends the perfect drug match with score, target & clear reasoning
-
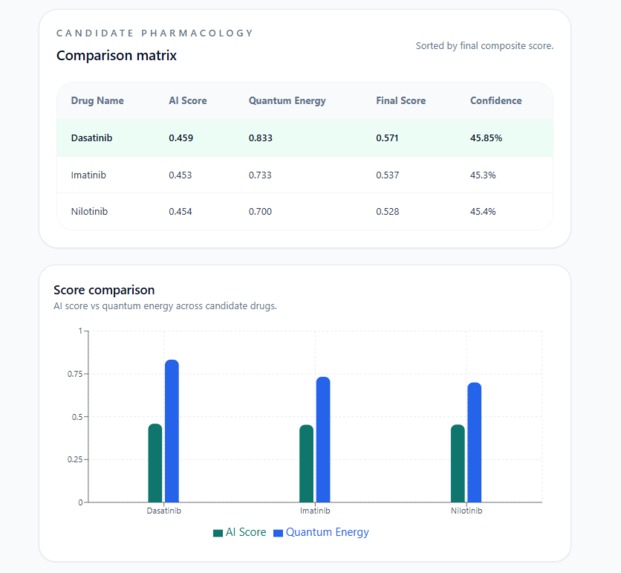
Comparison matrix ranks Dasatinib, Imatinib & Nilotinib by AI Score, Quantum Energy & Confidence the best drug match surfaced instantly.
-
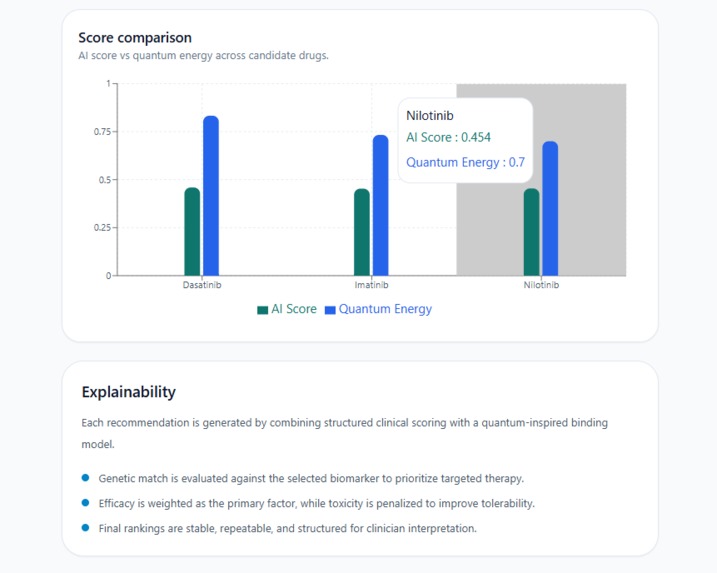
Interactive chart + Explainability panel hover for exact scores, see why every drug was ranked: genetic match, efficacy & toxicity balance.
Inspiration:-
It started with a question that bothered me more than it probably should have.
Why, in 2025, do cancer patients still go through months of chemotherapy with all the side effects, the exhaustion, the hope only to find out the drug didn't even work for their specific case?
I started reading about personalized medicine and kept hitting the same wall: every patient's cancer is genetically unique, but most treatment decisions are still made with population-level data. The bottleneck isn't a lack of drugs. There are hundreds of approved oncology drugs. The problem is the inability to quickly figure out which drug actually fits this patient's biology their specific disease type, their genetic markers, their protein targets.
A doctor today has to rely on clinical guidelines, experience, and trial-and-error. I wanted to build something that could give them a second opinion fast, explainable, and grounded in both AI scoring and molecular binding science.
That's where Q-BioTarget came from. And I built the whole thing solo.
What it does :-
Q-BioTarget is a clinical decision-support dashboard that matches a cancer patient's profile to the most suitable drug candidate with explainable reasoning a clinician can actually read and trust.
The flow is simple by design.
A clinician opens the Patient Intake panel and enters three things: a Patient ID, the disease type (for example, Blood Cancer), and the patient's genetic marker (for example, EGFR or BCR-ABL). They hit "Analyze patient profile" and the system takes it from there.
On the results side, the dashboard surfaces:
A Recommended Drug the top match with its name, indication, molecular target, final composite score, and confidence percentage. For patient P-003 with Blood Cancer and EGFR marker, the system recommended Dasatinib targeting BCR-ABL with a final score of 0.571 and 45.85% confidence.
A "Why this drug? explanation in plain English three bullet points that tell the clinician exactly why this drug was selected: balanced efficacy and genetic compatibility, quantum-inspired binding energy supporting stable target engagement, and toxicity minimization. No black box.
A Candidate Pharmacology comparison matrix a ranked table of all candidate drugs showing AI Score, Quantum Energy score, Final Score, and Confidence side by side. Every number is visible, every candidate is shown. The clinician can see why Dasatinib ranked above Imatinib and Nilotinib, not just that it did.
A Score comparison chart a Recharts bar chart plotting AI Score vs Quantum Energy for each candidate, with interactive tooltips on hover showing exact values per drug.
An Explainability section at the bottom that describes the scoring methodology in plain language: genetic match is evaluated against the selected biomarker, efficacy is weighted as the primary factor while toxicity is penalized, and final rankings are stable, repeatable, and structured for clinician interpretation.
The whole dashboard ends with a disclaimer I felt was important to include: "This system is a clinical decision-support prototype and should not replace professional medical judgment." That line matters. This is a tool to support doctors, not replace them.
How I built it :-
I kept the stack practical and the architecture clean.
The frontend is React with Tailwind CSS for styling and Recharts for the score comparison visualizations. Every component the patient intake form, the recommendation card, the comparison matrix, the explainability section was built to feel like something a clinician would actually be comfortable opening.
The backend is Python with FastAPI served through Uvicorn. Pydantic handles data validation on the incoming patient profile. The frontend and backend talk through a single analyze endpoint patient data goes in, ranked drug recommendations come out.
The core of the system is 'quantum_engine.py' my custom quantum-inspired binding simulation module. It takes the top drug candidates shortlisted by the AI scoring layer and computes a normalized binding energy score for each one. The scoring is deterministic and reproducible, inspired by quantum binding concepts like ground-state energy minimization and molecular affinity. It's not running on quantum hardware I want to be clear about that but the architecture is deliberately modular. The engine is a self-contained component; a real quantum backend using Qiskit Nature or PennyLane could slot in without touching anything else in the system.
The final score each drug receives is a composite of the AI Score and the Quantum Energy score, which is what gets ranked in the comparison matrix and drives the recommendation.
Drug data lives in 'drugs.json' a structured dataset I designed with fields mapping drug properties, indications, molecular targets, efficacy, and toxicity to mutation compatibility scores.
Tech stack :-
| Layer | Tech |
| Frontend | React + Tailwind CSS + Recharts |
| Backend | Python + FastAPI + Uvicorn |
| Validation | Pydantic |
| Simulation engine | Custom quantum_engine.py |
| Data | drugs.json |
Challenges I ran into :-
The hardest part wasn't the code. It was making the science honest and the output trustworthy as a solo builder, every decision landed entirely on me.
On the quantum side: I initially wanted to integrate an actual quantum SDK. But once I got into it, running a meaningful VQE simulation at prototype scale without misrepresenting what it's doing was deeper than the hackathon timeline allowed. So I made a deliberate call: build a transparent quantum-inspired simulation rather than a half-working Qiskit integration that looks impressive but produces meaningless output. I think that was the right call.
On the explainability side: The hardest design question wasn't "does the recommendation work?" it was "will a clinician trust it?" That's why I built the "Why this drug?" section and the full explainability panel. Every recommendation has to come with a reason, not just a score. Getting that language right clear enough for a non-technical clinician, honest enough to not overclaim went through several iterations.
On the UI side: Showing AI scores, quantum energy values, final composite scores, and confidence percentages all on one screen without overwhelming the user took real thought. The comparison matrix and the bar chart ended up being the key they let the clinician see the full picture while the recommendation card gives them the answer they actually came for.
On the technical side: CORS between React and FastAPI was the classic "works on my machine" problem. And designing drugs.json with scientifically reasonable fields not just made-up numbers required actual research into how drug-mutation compatibility and binding affinity are modelled in oncology.
Doing all of this alone meant there was no one to sanity-check decisions in real time. Every tradeoff was mine to make and live with.
Accomplishments that I'm proud of :-
The end-to-end pipeline working is the obvious one a patient ID, disease type, and genetic marker go in, and a ranked drug recommendation with full explainability comes out. That loop closing cleanly felt good, especially solo.
But the thing I'm most proud of is the explainability layer. It would have been easy to build a system that just spits out a drug name and a confidence score. Instead, every recommendation in Q-BioTarget comes with a plain-English explanation, a full candidate comparison, and a visible scoring breakdown. A clinician can look at the output and understand why not just what.
I'm also proud of how I handled the quantum piece. Calling it "quantum-inspired" and being upfront about what that means rather than dropping "Qiskit" in the README and hoping nobody asks felt like the more honest and interesting approach.
And the dashboard genuinely looks like a clinical product. Building something that feels this polished, end-to-end, alone, in a hackathon that's the part I'll remember.
What I learned :-
I learned a lot about molecular binding affinity what it actually means for a drug to fit a protein, why lower binding energy signals a stronger interaction, and where quantum computing has a genuine theoretical edge over classical simulation in this domain.
I learned that explainability is a design problem, not just a technical one. It's not enough to have a model that produces good outputs. The output has to be presented in a way that builds trust especially in a domain like oncology where the stakes are real.
I learned that building alone forces clarity. There's no one to hand off the confusing parts to. You either figure it out or it doesn't get built. That constraint made me a better decision-maker over the course of this hackathon.
And I learned that FastAPI is genuinely great. That one isn't deep, but it's true.
What's next for Q-BioTarget :-
The quantum engine is the most obvious upgrade. The architecture is already built for it quantum_engine.py is a clean, self-contained module. The next version replaces it with a real VQE simulation using Qiskit Nature or PennyLane, and eventually connects to a cloud QPU via IBM Quantum or AWS Braket. That's the version where the "quantum" in the name stops being inspired and starts being literal.
Beyond that :-
- Real genomic data connect to ClinVar and TCGA instead of my custom 'drugs.json', so the system works with actual patient mutation profiles
- Expanded drug coverage integrate with ChEMBL or PubChem for a clinically comprehensive drug library
- Clinical validation work with oncology researchers to validate binding energy predictions against known experimental results
- EHR integration connect Patient ID lookup to real electronic health record systems so clinicians don't have to manually enter what the system should already know
- Quantum vs classical benchmarking once the real quantum backend is in, systematically compare results against classical docking tools like AutoDock to measure where the approach actually wins
I built Q-BioTarget alone, over a single hackathon sprint. But it's a prototype with a real architecture, a real problem, a working demo, and a clear upgrade path.
That feels like something worth building on.



Log in or sign up for Devpost to join the conversation.