-
-
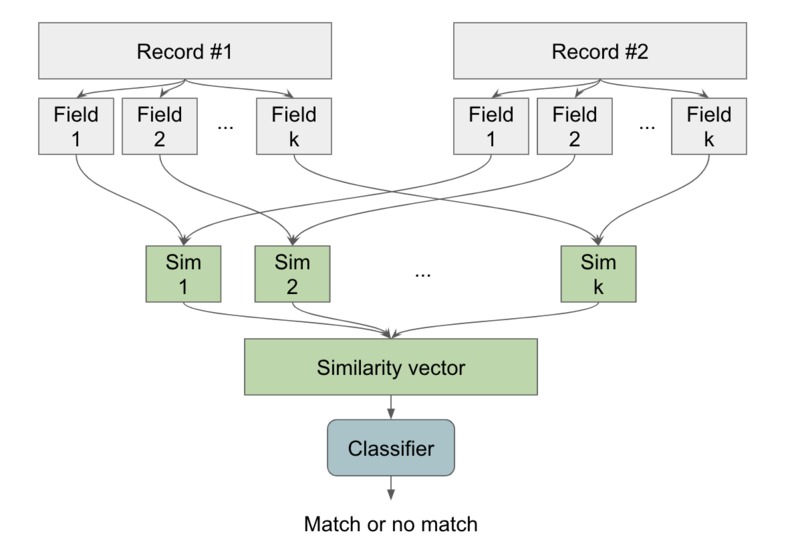
High-level idea of novel solution. We compute the similarity between each field, and use all of these to decide if the records match.
-
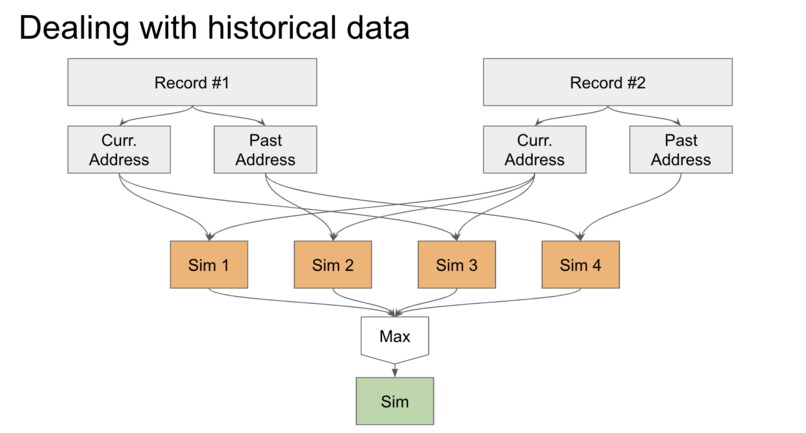
Novel solution to deal with historical data. We may miss matches if we only compare corresponding fields, so we take the max over all pairs.
-
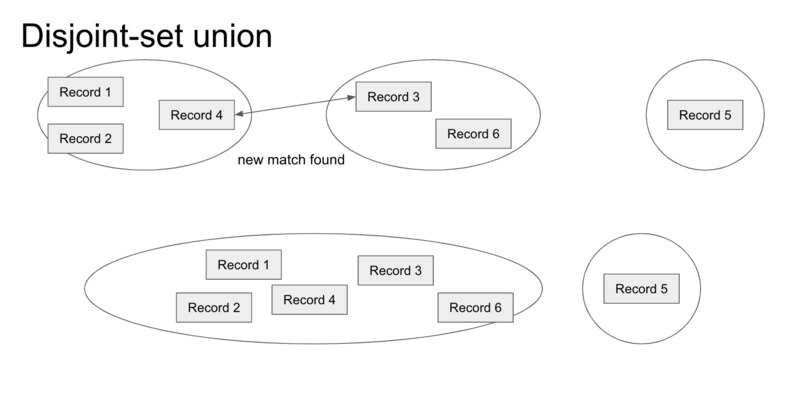
DSU gives an online solution to merging groups of patient records once we find matches between individual records.
-
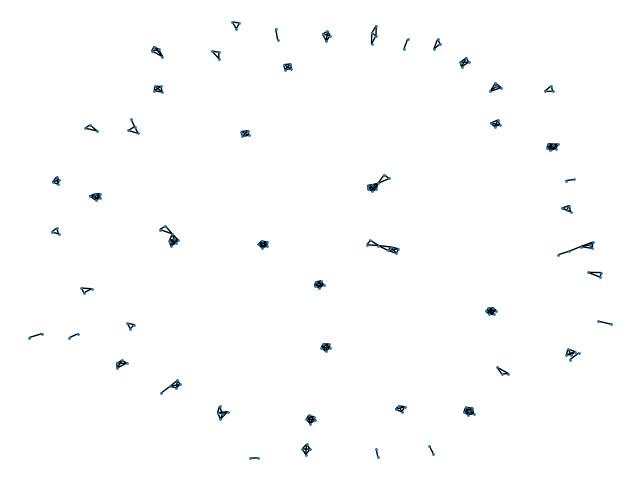

Matchings found using novel solution in the test data. Edges are predicted matchings between records, and connected components are patients.
Inspiration
We were inspired by the first Office Ally challenge at LA Hacks 2020! When this challenge was presented, we couldn't believe that such an important problem was unsolved. Just based on intuition, it seemed relatively easy to judge if two patient profiles are the same. Upon further thought and after examining Office Ally's example data, we realized how difficult it was to account for all of the edge cases in this problem, and we were very motivated to come up with novel and effective solutions.
What it does
The patient matching problem is to find all patient records that belong to the same person in a large dataset of patient records. We explored two different solutions to this problem by implementing an algorithm inspired by Office Ally's POC, and creating a novel algorithm.
How it works
Implementing Office Ally's POC Algorithm
Inspired by Office Ally's POC, we created a similar algorithm in Python with additional features and heuristics to improve the accuracy of patient matching. Given that the patient records come from different sources with different formats, it was important to first clean the data and standardize the fields. We utilized the Pandas library to tidy the data so that all strings fields were upper-case/alphanumeric strings, missing values were filled in (with 0 or empty strings), and dates and zip codes were formatted properly.
Next, our algorithm computed SHA-1 partial hashes for every patient record so that accurate records would map to the same partial hash for the initial matching. We hashed the first name (first 3 letters only), last name (first 3 letters only), date of birth, and gender since these pieces of information together are (essentially) unique identifiers with low collision rates. For good records, this means that we can quickly group all entries together that have the same partial hash and identify the initial matchings for patient records.
However, patient records often contain missing, mis-typed, or duplicated information, and these complications could endanger lives and cost billions in the healthcare industry. To improve patient matching, our algorithm uses heuristics to determine when two patient groups are similar enough to be the same. For each pair of initial patient groupings, we estimate a similarity score based on Soundex tokens/Levenshtein distances for first and last name, and confidence in similarities of birthdate, gender, street addresses, states, etc. Our heuristics were guided by our intuition: for example, we decided that two patient groups having the same birthdate AND similar names/street address was a much stronger indicator of a match than if they were just the same gender with similar names.
Lastly, our algorithm uses an optimized implementation of disjoint-set union to identify the final patient matchings in an output CSV file and SQL table. Overall, our implementation of the initial POC algorithm performed well. Compared to the "true" 65 group ID matchings for the 201 patients, our algorithm found 67 different patients, with ~6 false positives and ~10 false negatives.
Creating a Novel Algorithm
We claim that we can reduce the problem of finding all matchings to a decision problem: given two patient records, decide whether they belong to the same patient. This is because if we can solve this decision problem, we can use it to build an undirected graph where nodes are records and there are edges between records belonging to the same patient. Then, a connected component in this graph corresponds to all of the records belonging to one patient.
As an overview, our novel algorithm to solve this decision problem is to estimate how similar each corresponding field in the records are to create a similarity vector, and then use this similarity vector to decide whether they are the same.
To compute the similarity between each field in the records, we handcrafted models.
Before anything was done, we attempted to correct the (city, zipcode, state) tuple. Since all three are related, we can try and fill in any missing information. Going through the test set, we also noticed that someone had an IATA code as their city, so we found a list of all US airport IATA code and their corresponding cities so we can switch from IATA code to a string of a city name. Using the zipcodes modules in python, we get the zipcode information from zipcode and city, if they exist. If only a single field is missing or wrong (typically zipcode or misspelled states), we look to see if there are zipcode object that matches the two other fields and if so, update it accordingly—it is exceedingly rare for there to be duplicates at this point.
We thought about filling perhaps two blanks if we are only given one field (like city or zipcode). However, there are many duplicate cities in different states, and zipcode could be typed incorrectly. Thus, we chose to ignore such cases and leave them as is.
The next thing we did is to lowercase everything. Having a consistent case will help us with the string distance in the future. Levenshtein string distance plays a great role in our algorithm.
For names, we mainly used Soundex and Levenshtein. Soundex is great for names that sound similar— katy Vs katie. We get the Soundex representation and look at the Levenshtein distance. However, this doesn’t cover everything. Seeing the number of typos within the training set, it is possible that someone types two characters incorrectly and thus Soundex is no longer effective. For this reason, we also use Levenshtein on the entire name. We then return the max similarity between Soundex distance and Levenshtein distance.
This resulted in a name similarity model that captures names that are generally similar in length. What it failed to capture, however, is nicknames. For example, Kim and Kimberly would not result in a match despite it clearly being one. To combats this, we found a csv file of all the common nicknames. From the two names that we are given, we generate two lists of similar names. We do the Soundex + Levenshtein on all possible pairs and return the highest similarity.
For Date of Birth, Patient Acct #, and zip code, we simply looked at the Levenshtein distance.
For streets, we elected to keep street 1 and street 2 as separate fields. This is because going through the training set, we saw many incomplete street 2’s. Next, we found a list of street name abbreviation and expanded all abbreviations.
We had two different methods to test similarities between street names. The first is Levenshtein once again. This would account for any misspellings in the form, for example, 59 Sumer Ridge Court versus 59 Summer Ridge Court The other method is to combat people filling in the form but leaving words out, for example,485 Hanson St. versus 485 Hanson Street East. For each unique word in the two street strings, we have a separate character than represents them. We then get a hash of each street string and use Levenshtein once again. (Side note: we should have used some other metric to determine if a word is truly unique or another typo, but unfortunately we ran out of time). We return the max similarity between using hashes and using plain Levenshtein.
For gender, we mapped F, M to female and male, and everything else was treated as an empty string. <we should prolly use strict Levenshtein here!! Femole vs female for example).
For states, it was directly checking if they are the same or if fields are missing. All states are converted to lowercased abbreviations.
To decide whether a similarity vector comes from two records from the same patient, we used logistic regression. Our motivation for this model choice was we wanted a model with low model capacity and high generalization so we can fit the small training set that we are given without overfitting. We considered other models like multilayer perceptrons and gradient-boosted decision trees, but ultimately concluded that these models were much too complicated given our training set available.
In our hold-out validation testing, we trained on the first 150 rows of the given dataset and tested on the last 50 rows. Our novel algorithm achieved a precision of 0.93, a recall of 1.0, and a F1 score of 0.97, with 4 false positives and no false negatives.
More Details
In this section, we discuss some interesting implementation details of our models that required ingenuity.
For our novel algorithm, we needed to think about how to deal with records containing both current and past data. For example, a patient can move and change addresses, so the current address fields may not be the same, but the current address field for one record and the and past address field for the second are the same. If we were to compute similarity for corresponding records only, we would miss this interaction. Our solution was to notice that if we look at all possible pairings of the data: (past, past), (past, current), (current, past), and (current, current), if any of these fields are very similar, then this field should have high similarity. Our final solution is to take the max over all pairwise similarities computed for fields with historical data and use this as the similarity score.
For both approaches, we needed a way to fill out Group IDs for the patients after we've found the pairwise matchings. As we discussed above, this is equivalent to finding the connected components in the induced graph and giving one Group ID per connected component. Performing a graph search at the end of the algorithm is an offline solution to this problem, but it requires another pass through the entire graph, and for us to store the entire graph in memory. We instead propose an online solution using a disjoint-set data structure. We initialize each record to its own set, and whenever we find a matching between two records, we use the disjoint-set data structure to union the two records and their corresponding sets together. Our optimized implementation of disjoint-set union can perform this operation in near O(1) time. Our final output is an identifier for each record, and records in the same group have the same identifier, as desired.
Challenges
We enjoyed the challenge of working with a small data set. While we had to pivot from our initial idea of using deep learning, the small sample set allowed us to to better see the inconsistencies and understand the edge cases. Another challenge was figuring out how to deduplicate the patient matchings once we determined similarity between two patient groups. Learning about the disjoint-set data structure and applying it to our project was a fun challenge.
Accomplishments
While we were initially intimidated by the lengthy POC algorithm given to us, we are proud that we were ultimately able to understand and implement parts of it with improvements. We are also excited about how we designed and implemented our novel approach in such a short period of time.
Learnings
Like most datasets in the real world, the sample records that we were given were messy and inconsistent. We felt like this experience was a valuable exercise in cleaning datasets and helped us realize the importance of standardization. We also learned more about the challenges faced in the healthcare industry and how creative solutions could make tremendous financial and social impacts.
What's next for Patiently
Since we constructed our heuristics and model for both of our algorithms without bias, we expect Patiently's solutions to perform well on other patient record test sets. However, having much larger set of data could potentially allow us to explore additional solutions as well using machine learning techniques.
In order to solve the patient matching problem that the healthcare industry faces, an effective solution needs to be secure and scalable. Patiently would work on refining our algorithms with these criteria in mind, while still retaining the high accuracy.
Github: https://github.com/helenawu1998/officeally_challenge




Log in or sign up for Devpost to join the conversation.