-
-
Home section
-

Vision Section
-
How it works section
-
Ethics section
-
Report - Bio section 1
-

Report - Bio section 2
-
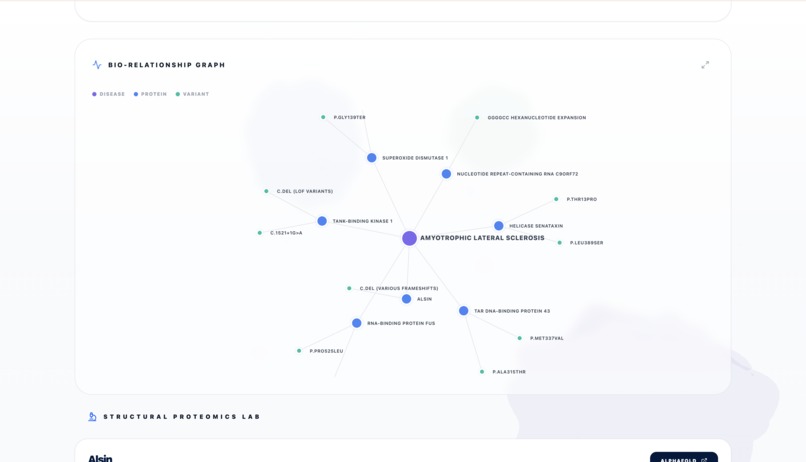
Report - Bio section Graph connection
-
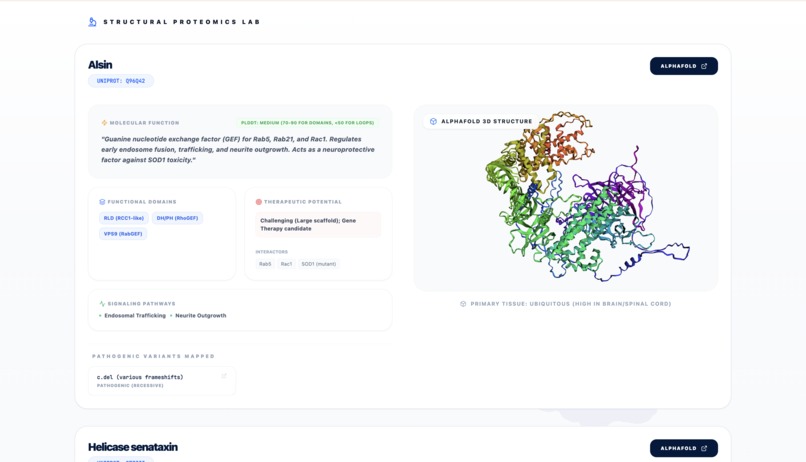
Report - Bio section - Proteic Lab
-
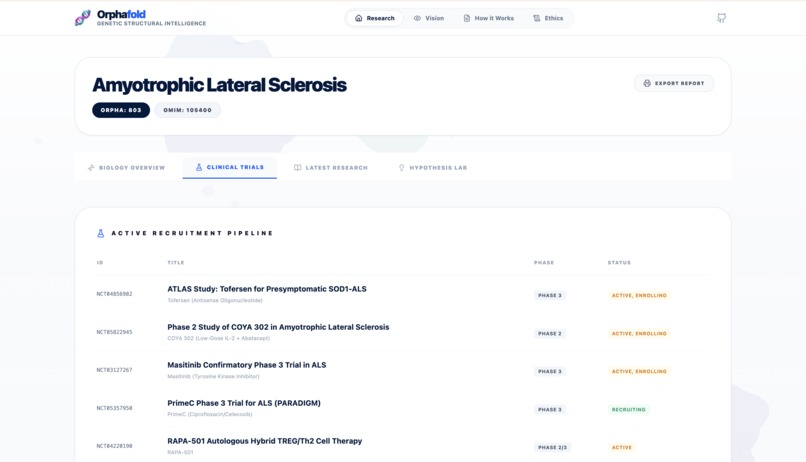
Report - Clinical Trial section 1
-
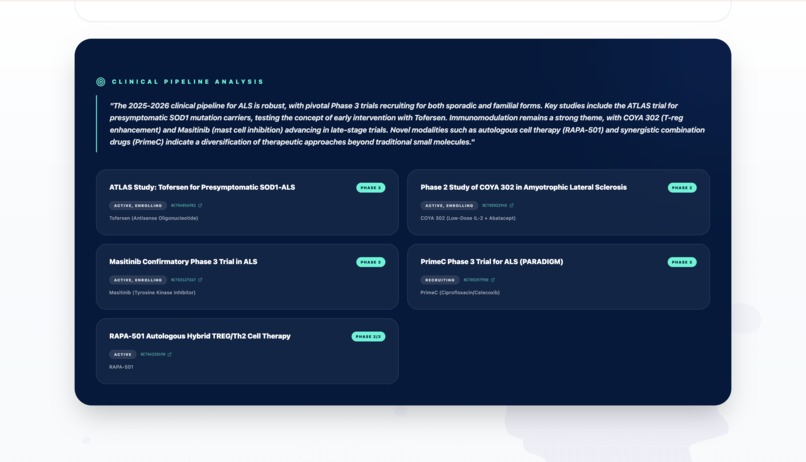
Report - Clinical Trial section 2
-
Report - Bibliography
-
Report - Hypothesis Lab 1
-

Report - Hypothesis Lab 2
Project Story
Inspiration
My name is Paul, and I’m a computer scientist dedicated to AI for Good. My journey in this field reached a milestone during the Paris 2024 Olympic Games, where I developed Prometheus BCI—a system that allowed people with severe motor disabilities to carry the Olympic flame using only their brainwaves. As a co-founder of the neurotech company Inclusive Brains, we released this technology as open source to push boundaries for inclusion.
However, the true "Why" behind OrphaFold is deeper. I have spent years working alongside people suffering from neurodegenerative diseases that eventually led to total tetraplegia. Seeing their daily battle, I witnessed first-hand the devastating speed of these conditions versus the slow pace of drug discovery. I often asked myself: How can Generative AI act as a catalyst for innovation where traditional research stalls?
OrphaFold is my attempt to answer that. Today, 300 million people live with a rare disease, yet 95% of these 7,000+ conditions have no approved treatment. Research is often hindered by fragmented data—clinical profiles are in one silo (Orphanet), molecular data in another (UniProt), and structural insights (AlphaFold) remain disconnected from the literature (PubMed). OrphaFold was inspired by the need to bridge these silos using Structural Intelligence. We wanted to see if an agentic system could mirror the workflow of a high-level geneticist, synthesizing complex biological data to identify drug repurposing opportunities that human researchers might miss.
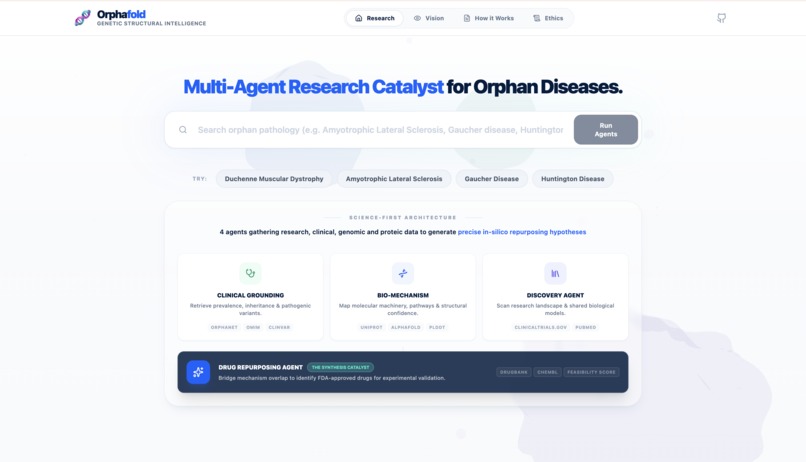
What it does
OrphaFold is an agentic research platform that automates the discovery phase for orphan diseases. It orchestrates a pipeline of 4 specialized Gemini 3 agents:
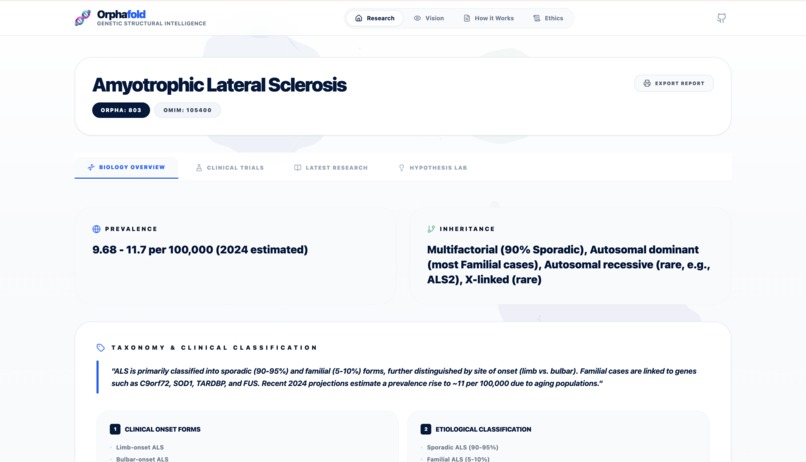
- Clinical Grounding: Establishes the baseline using Orphanet and OMIM APIs.
- Bio-Mechanism: Analyzes the molecular pathophysiology through Alphafold DB, UniProt and ClinVar.
- Discovery Agent: Synthesizes the latest research from PubMed and ClinicalTrials.gov.
- Hypothesis Lab: The "brain" that performs structural homology analysis and binding pocket comparison to propose therapeutic candidates by bridging mechanism overlap.
The system generates a full structural report including interactive 3D protein visualizations and a comparative discovery analysis between different diseases.
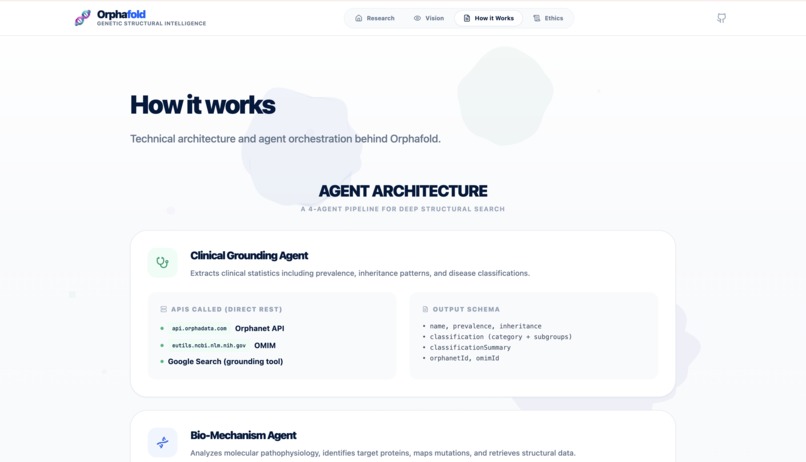
How we built it
The core of OrphaFold is a Multi-Agent Orchestration layer built on Gemini 3.
- The Engine: We utilized Gemini’s deep reasoning and large context window to manage parallel API enrichment from Alphafold DB, Orphanet, UniProt, and NCBI.
- The Frontend: Built with React/Vite, we implemented a custom Structural Proteomics Lab using 3D rendering for AlphaFold pdb models.
- Orchestrated Agentic Reasoning: We designed a multi-layered workflow where agents autonomously cross-validate clinical registries with structural proteomic data, transforming raw information into synthesized discovery paths. For example, the system correlates pathogenic variants from ClinVar with specific functional domains in UniProt to assess candidate confidence based on pLDDT and structural significance.
Gemini Integration Summary
OrphaFold utilizes Gemini 3 as the core reasoning engine for its multi-agent architecture. The application is built around four specialized agents that collaborate to analyze rare diseases:
- Clinical Grounding: Uses Search Grounding to verify prevalence and inheritance patterns from trusted medical sources.
- Bio-Mechanism: Reasons over raw UniProt and NCBI data to map molecular pathways and druggability scores.
- Discovery: Synthesizes literature and identifies ongoing clinical trials via PubMed.
- Hypothesis Lab (Repurposing): Acts as the synthesis catalyst, using Gemini’s large context window to bridge disease mechanisms with existing drug properties through structural homology analysis and binding pocket comparison.
Gemini is central to our project’s success, enabling us to perform complex multi-step reasoning over structured biological data and unstructured research papers simultaneously. We specifically leverage Gemini’s ability to maintain high-fidelity context across multiple specialized agent prompts, allowing for a 'synthetic researcher' experience. By integrating Gemini's reasoning with structural data from AlphaFold, we create a tool that moves beyond simple search into predictive discovery, optimized for the precision and low-latency requirements of modern scientific research.
Challenges we ran into
The biggest challenge was Data Synthesis latency. Orchestrating 4 agents while querying multiple REST APIs simultaneously requires precise prompt engineering to prevent "hallucinations" in a medical context. We solved this by implementing a "Pre-Flight" layer that fetches raw structural data first, feeding it into Gemini as a grounded context, ensuring that every hypothesis is backed by a verified UniProt ID or PubMed citation.
Accomplishments that we're proud of
- Structural Homology Hypotheses: We are proud of the "Hypothesis Lab" which can identify that two very different diseases might share a similar structural defect, potentially opening the door for shared drug treatments.
- Seamless Integration: Mapping 3D AlphaFold structures directly to real-time clinical trial data in a single dashboard.
- Researcher-Ready Exports: Implementing the RIS export feature, allowing researchers to instantly save synthesized bibliographies into Zotero or Mendeley.
What we learned
We learned that Gemini 3’s reasoning capabilities are a game-changer for scientific research !! Unlike previous models, Gemini 3 can follow complex biological "logic chains" (e.g., "If mutation X affects domain Y, then drug Z which targets that domain might work"). We also realized the power of agentic workflows to handle the "cold start" problem in rare disease research, where data is extremely scarce.
What's next for OrphaFold
Our next step is to integrate direct docking simulations. We want to transform our hypotheses into predictive scores by running in-silico simulations directly within the agentic loop. We are also exploring a transition to the Agent Development Kit (ADK) framework to leverage its more comprehensive orchestration capabilities.
Winning this hackathon would provide the necessary visibility to launch pilot beta tests with geneticists and rare disease researchers, accelerating our journey from discovery to impact.
Built With
- agentic-ai
- alphafold
- clinvar
- gemini
- google-ai-studio
- google-genai-sdk
- javascript
- orphanet
- pubmed
- react-native















Log in or sign up for Devpost to join the conversation.