-
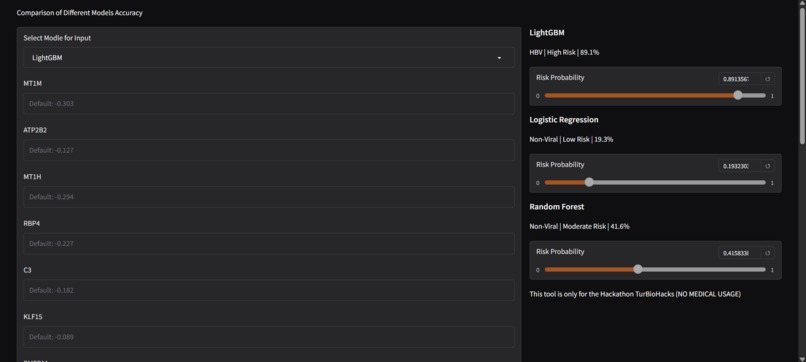
A demonstration of all 3 methods (Linear Regression, Random Forest, and LightGBM) predicting HBV-related HCC.
-
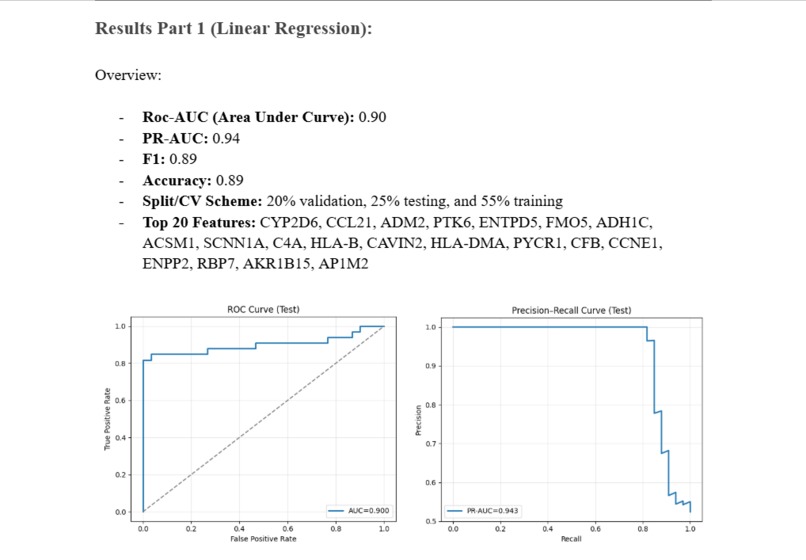
A quick overview of the accuracy of the Linear Regression Model
-
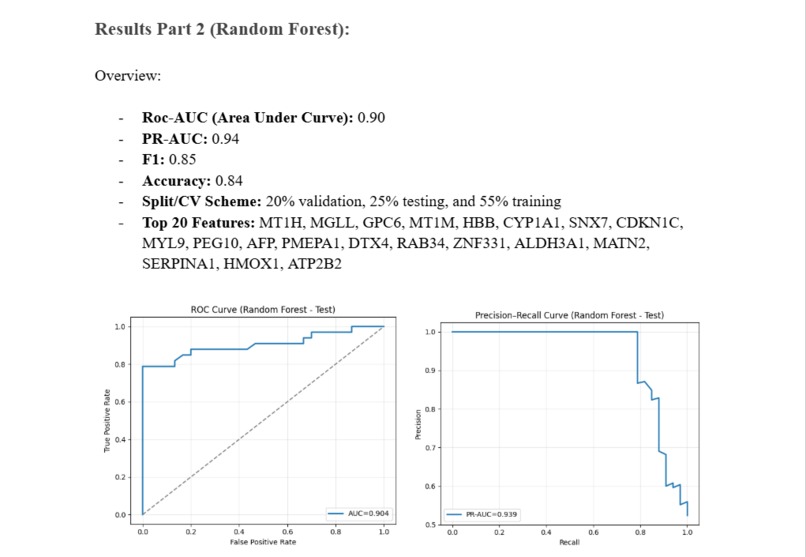
A quick overview of the accuracy of the Random Forest Model
-
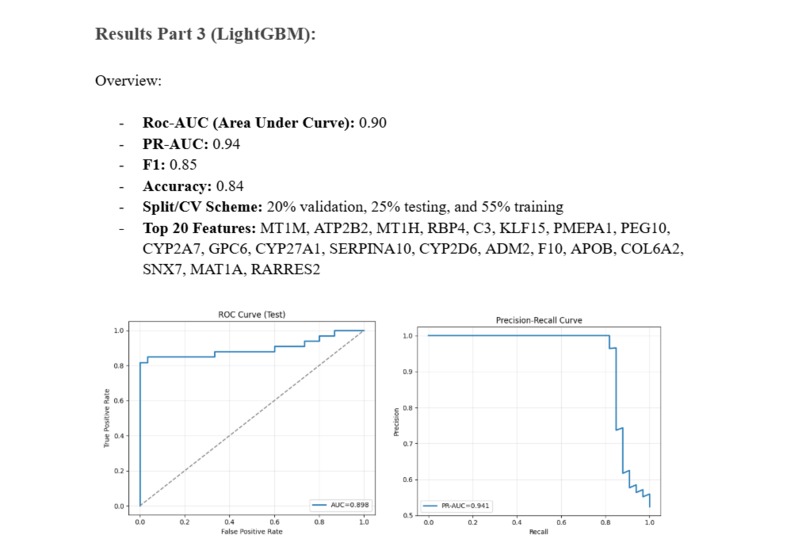
A quick overview of the accuracy of the LightGBM Model
-
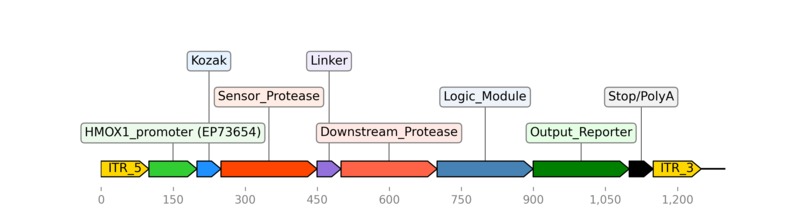
A hypothetical genetic circuit using the HMOX1 promoter (EP73654) to regulate gene expression.
-
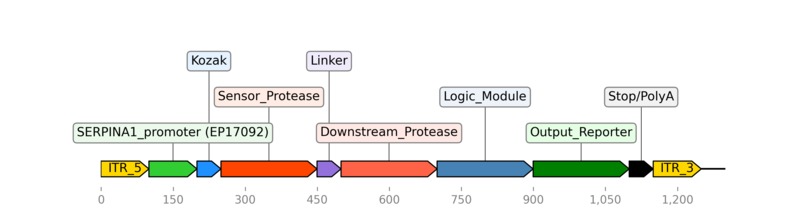
A hypothetical genetic circuit using the SERPINA1 promoter (EP17092) to regulate gene expression.
This will be a brief description about the project as the majority of the work can be found in the Google Doc link provided below :)
Inspiration
This project is mainly inspired by the Problem Packet and the Postcard Method by Dr. Endy. This project comes as a combination of the following "postcards":
**- The rise of automation (In Factories, Programming...)**
**- The availability of public data (Not only in Biology)**
**- STEM cells and being able to modify DNA to cure diseases (like Sickle Cell)**
This is basically my attempt at combining all these different postcards with the problem statement into one hackathon project.
What it does
This project does two essential steps:
1) Classify HBV vs Non-Viral HCC: This project employs three different strategies (Linear Regression, Random Forest, and LightGBM) in order to classify HBV-related HCC using expression levels. It also provides a friendly user interface for you to compare the accuracy and differences between the different methods.
2) Create a genetic circuit: After identifying the top features (biomarkers), the project can also search for promoters that react to those biomarkers and compile a hypothetical genetic circuit to detect HBV cancer (basically programming a cell to be able to identify HBV-related HCC in the body).
How we built it
The project was programmed entirely in Python using publicly available datasets such as TCGA-LIHC and The Eukaryotic Promoter Database (EPD). All attributions for any software used can be found in the attached Google Doc and/or in the code base.
Results (Quick Overview)
Note: The Random Forest method was the best performing (practical applications).
Results Part 1 (Random Forest): _ Best Performing One In Terms of Biological Significance_
Roc-AUC (Area Under Curve): 0.90 PR-AUC: 0.94 F1: 0.85 Accuracy: 0.84 Split/CV Scheme: 20% validation, 25% testing, and 55% training Top 20 Features: MT1H, MGLL, GPC6, MT1M, HBB, CYP1A1, SNX7, CDKN1C, MYL9, PEG10, AFP, PMEPA1, DTX4, RAB34, ZNF331, ALDH3A1, MATN2, SERPINA1, HMOX1, ATP2B2
Results Part 2 (Linear Regression):
Roc-AUC (Area Under Curve): 0.90 PR-AUC: 0.94 F1: 0.89 Accuracy: 0.89 Split/CV Scheme: 20% validation, 25% testing, and 55% training Top 20 Features: CYP2D6, CCL21, ADM2, PTK6, ENTPD5, FMO5, ADH1C, ACSM1, SCNN1A, C4A, HLA-B, CAVIN2, HLA-DMA, PYCR1, CFB, CCNE1, ENPP2, RBP7, AKR1B15, AP1M2
Results Part 3 (LightGBM):
Roc-AUC (Area Under Curve): 0.90 PR-AUC: 0.94 F1: 0.85 Accuracy: 0.84 Split/CV Scheme: 20% validation, 25% testing, and 55% training Top 20 Features: MT1M, ATP2B2, MT1H, RBP4, C3, KLF15, PMEPA1, PEG10, CYP2A7, GPC6, CYP27A1, SERPINA10, CYP2D6, ADM2, F10, APOB, COL6A2, SNX7, MAT1A, RARRES2
More detailed information (including charts) is in the attached document.
What we learned
This project taught me a lot. Specifically, the Random Forest method was new for me to experiment with. I also feel like the actual issue that we are trying to solve here (HCC) has become a lot clearer to me, and it introduced something that I might be interested in working on (even outside the hackathon).
What's next for Oncophoros
There is a lot more to come for the future of Oncophoros. One really interesting application would be to use generative AI to design a promoter specific to each biomarker, allowing for more accurate detection. Additionally, further work can be completed on the generation of genetic circuits (through the combination of in-silico and in-vitro lab methods, it could be possible to generate plausible genetic circuits to be used in labs and, in the future, aid in the design of medical smart therapeutics to treat cancer).







Log in or sign up for Devpost to join the conversation.