-
-
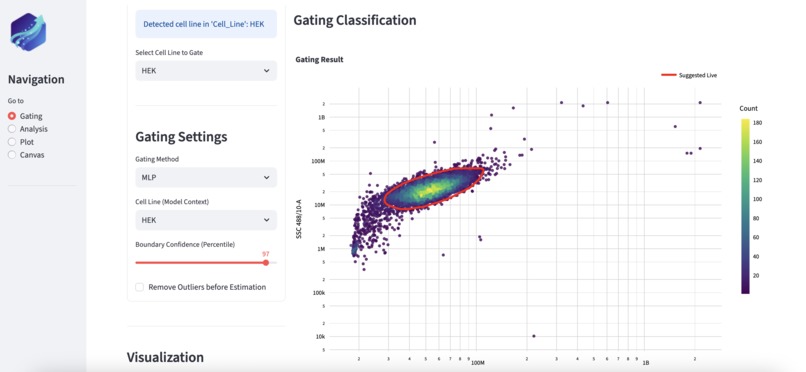
Automatic live cell gating with Glow. The decision boundary as well as cell population distributions are clearly visualized.
-
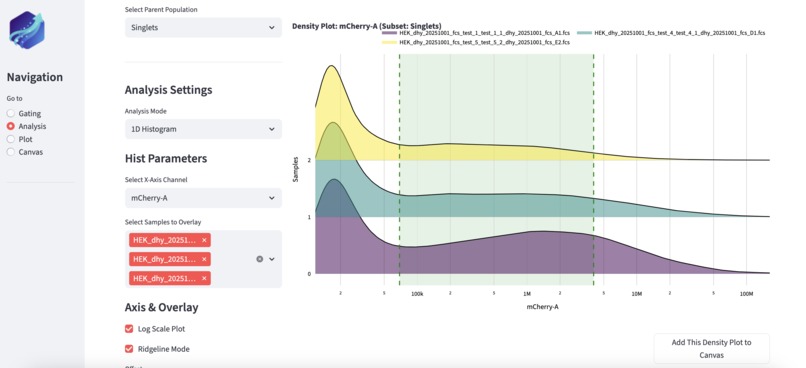
User can define custom fluorophore gating based on stacked comparison of flow data.
-
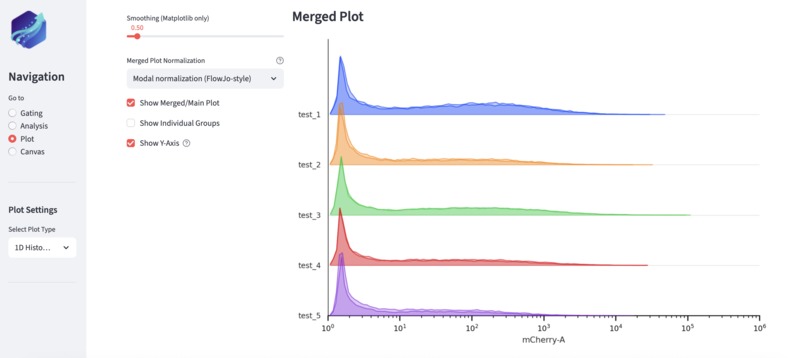
1D histogram of fluorophore channels generated by Glow. Data can be easily comparied between individual experiments.
-
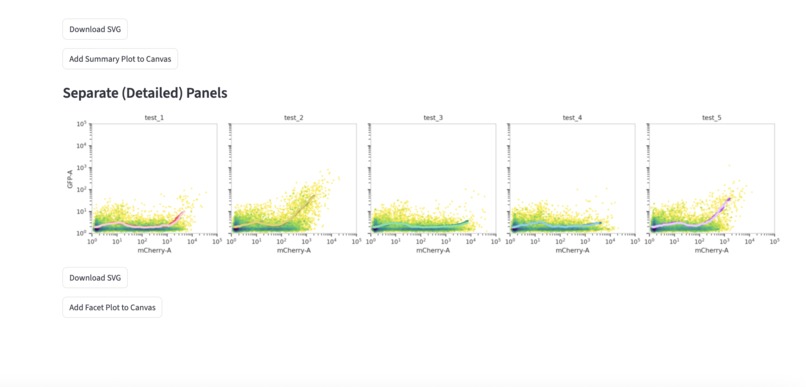
2d density plots with rolling means for comparisons. Data distributions can be clearly compared between individual experimental conditions.
-
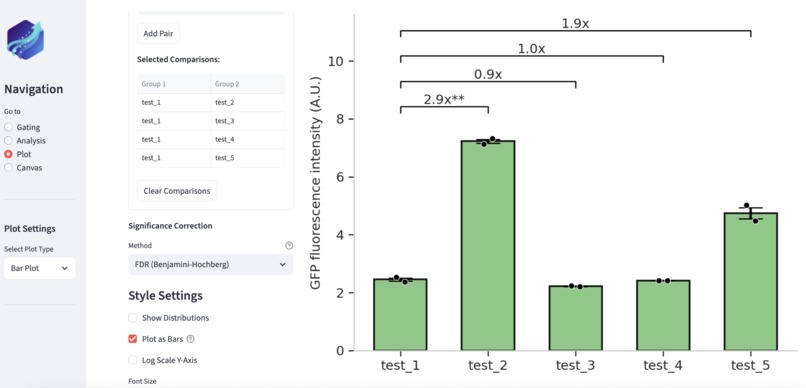
Glow can generate publication-level plots, with built-in comparison and statistic tests tailed to user's needs.
-
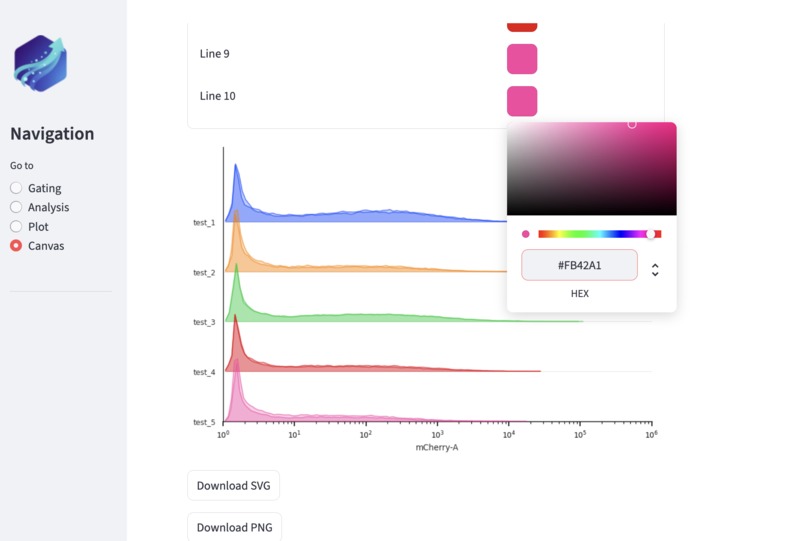
User can easily customize plots on canvas -- including font size, figure size, and colors.
Inspiration
Flow cytometry is a cornerstone of biological research, but the downstream analysis, specifically gating, remains a significant bottleneck. Traditional manual gating is time-consuming, subjective, and difficult to reproduce across large datasets, while existing automated solutions often lack the flexibility to adapt to specific experimental conditions or cell lines. We wanted to bridge this gap by building a tool that combines the intuitive control of manual gating with the speed and consistency of machine learning. Our goal was to create a system where the software can actively learns from the researcher's expertise to improve over time.
What it does
Glow is an end-to-end flow cytometry analysis pipeline wrapped in a user-friendly, interactive interface. It streamlines the entire workflow from raw data to publication-ready figures:
Automated & Interactive Gating: It automatically identifies Live/Dead populations and discriminates Singlets/Doublets using various algorithms (MLP, GMM, Density-based, and DBSCAN). Users can accept these suggestions or manually override them using interactive polygon tools.
Active Learning: The core innovation of Glow is its ability to learn from user feedback. When a user manually corrects a gate, the system logs this input to refine its future suggestions for that specific cell line.
Metadata Management: It simplifies the ingestion of metadata, allowing for auto-detection from filenames or manual upload via spreadsheets, ensuring experimental conditions are tracked accurately.
Advanced Analysis & Visualization: Beyond standard gating, it calculates transfection efficiencies, supports 2D quadratic gating, and generates high-quality visualizations (1D Stack Plots, 2D Running Means) that can be customized on a dedicated canvas and exported as SVGs.
How we built it
We built Glow using a modular approach to ensure cross-platform compatibility and scalability. We used Google Gemini 3 Pro API to develop this platform end-to-end.
Backend: The core logic is written in Python, utilizing libraries like scikit-learn for the machine learning components (Gaussian Mixture Models, MLPs, DBSCAN) and standard flow cytometry libraries for parsing .fcs files.
Frontend: We used Streamlit to build a reactive web-based interface that allows for real-time plotting and interaction without requiring users to write code.
Machine Learning Pipeline: We implemented an Active Learning loop where the "accept/reject" actions in the UI serve as labeling events, dynamically updating the model's understanding of specific cell populations.
Packaging: To ensure accessibility for non-coders, we packaged the application into standalone executables for Windows and Mac, removing the need for command-line installation or environment management on those platforms.
Development Acceleration: We utilized Gemini to accelerate our development process, using it to help optimize our gating algorithms and streamline the UI code structure.
Challenges we ran into
Balancing Automation with Control: Designing a UI that offers powerful automated suggestions without taking control away from the user was difficult. We had to ensure the "Manual Gating" override was fluid and that the system correctly interpreted these overrides as training data.
Handling Data Variability: Flow cytometry data can vary significantly between machines and experimental days. Tuning the default parameters of machine learning models for each cell line to work broadly across different datasets while remaining specific enough to be useful was a major challenge.
Interactive Plotting Performance: Rendering thousands of data points in a browser-based interface (Streamlit) while maintaining a responsive experience during manual gating required careful optimization of our plotting libraries and downsampling of the data.
Accomplishments that we're proud of
The "Canvas" Feature: We are particularly proud of the Canvas customization tool. It solves a major pain point for researchers by allowing them to tweak plot aesthetics (fonts, colors) within the app and export directly to SVG for Illustrator, saving hours of post-processing time.
True Cross-Platform Support: Successfully compiling the Python environment into standalone apps for Windows and Mac. This makes the tool immediately accessible to biologists who may not be comfortable with Linux or command-line interfaces.
Seamless Active Learning: Implementing a working feedback loop where the tool actually gets "smarter" the more you use it, rather than just being a static set of hard-coded rules.
What we learned
The Power of AI-Assisted Development: We were surprised by just how powerful the Gemini agent was as a force multiplier for our team. It didn't just help with syntax; it acted as a tireless pair programmer, accelerating our ability to debug complex Streamlit state issues, refactor our machine learning pipeline, and rapidly prototype the UI. This allowed us to focus on the biological logic rather than getting stuck on implementation details.
Synergy in Interdisciplinary Teamwork: We learned that building a tool for biologists requires more than just coding skills—it demands constant translation between computational logic and experimental reality. Our team's success came from blending our distinct strengths: validating the algorithms against real biological use cases while ensuring the software architecture remained robust and scalable.
User Experience is Critical: Even the best algorithm is useless if the interface is clunky. We learned that abstracting away the complexity of the underlying ML models and providing a responsive, intuitive "canvas" was just as important as the accuracy of the gating itself.
What's next for Glow
Batch Processing: Improving the backend to handle batch processing of hundreds of files simultaneously for high-throughput screening.
Cloud Deployment: Moving the processing pipeline to the cloud to allow users to analyze massive datasets without being limited by their local machine's RAM.






Log in or sign up for Devpost to join the conversation.