-
-
GLIMPSE user authentication.
-
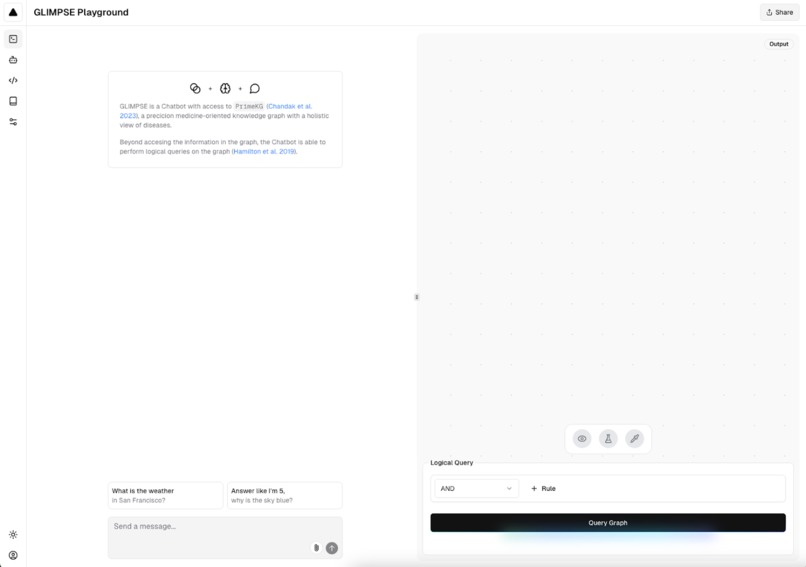
Example of GLIMPSE chat and biomedical graph visualization interface.
-
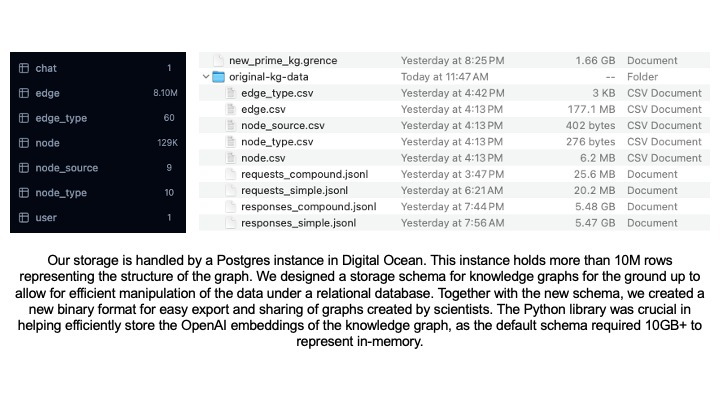
Efficient algorithms to enable knowledge graph and embedding storage with over 10M rows.
-
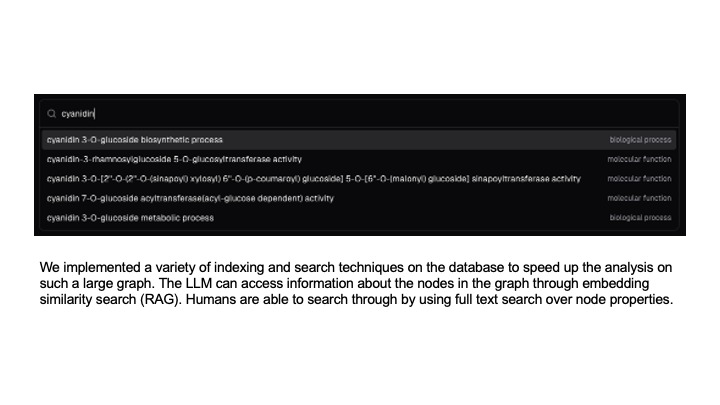
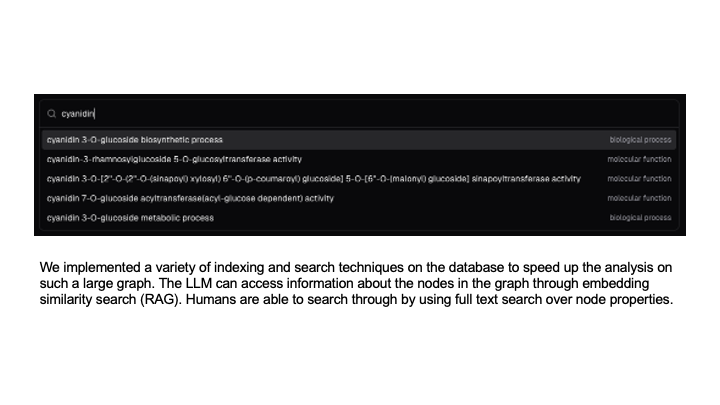
Indexing and searching techniques for GQE query construction.
-
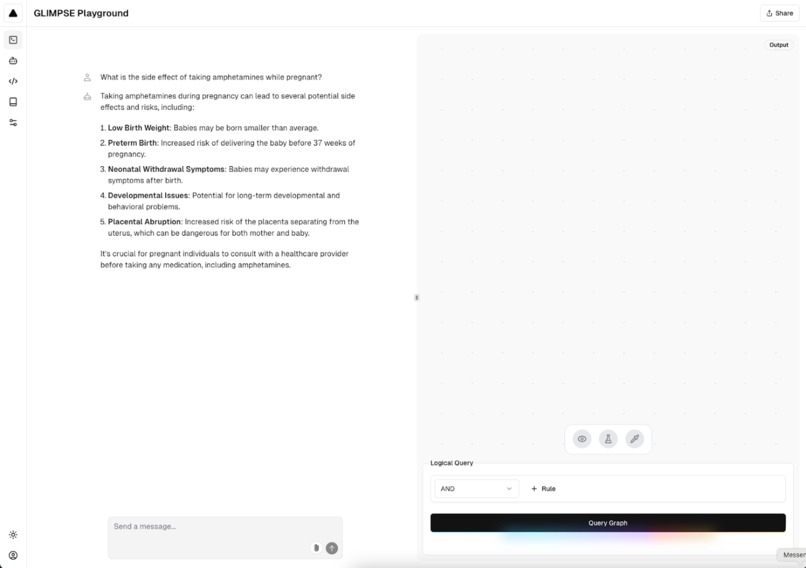
Example of GLIMPSE query in light mode.
-
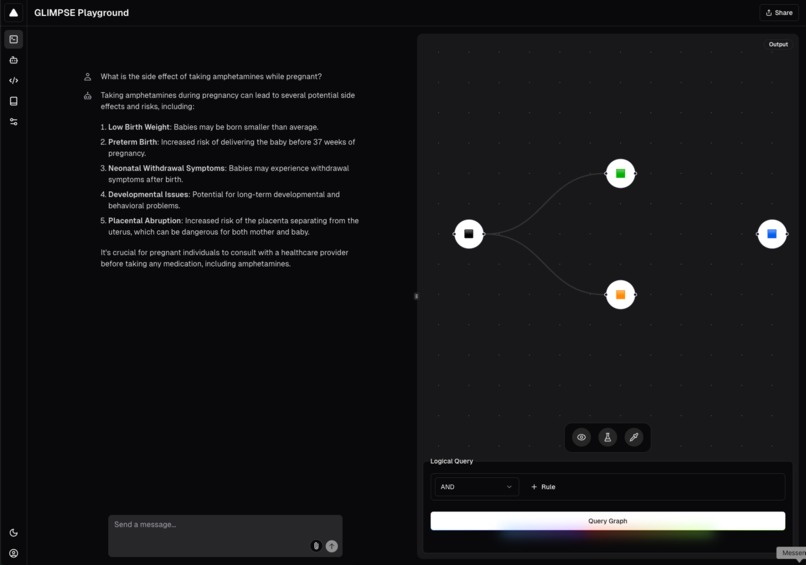
Example of GLIMPSE query in dark mode.
-
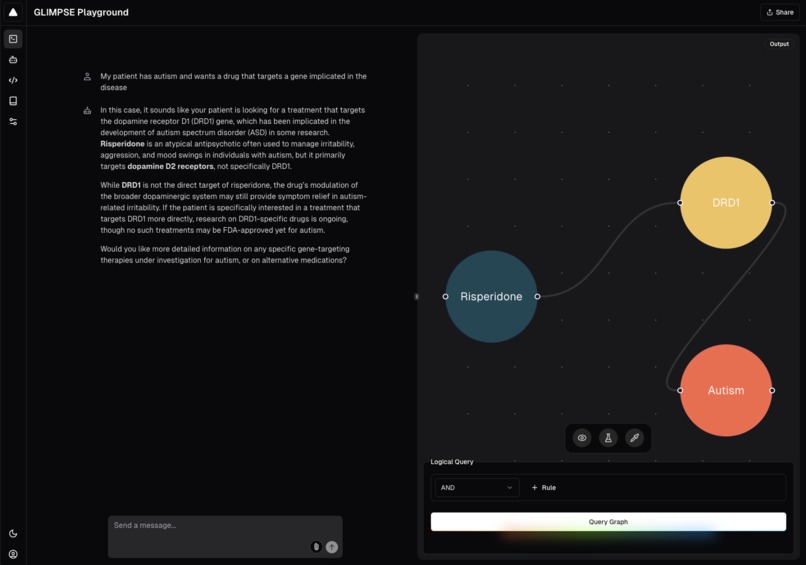
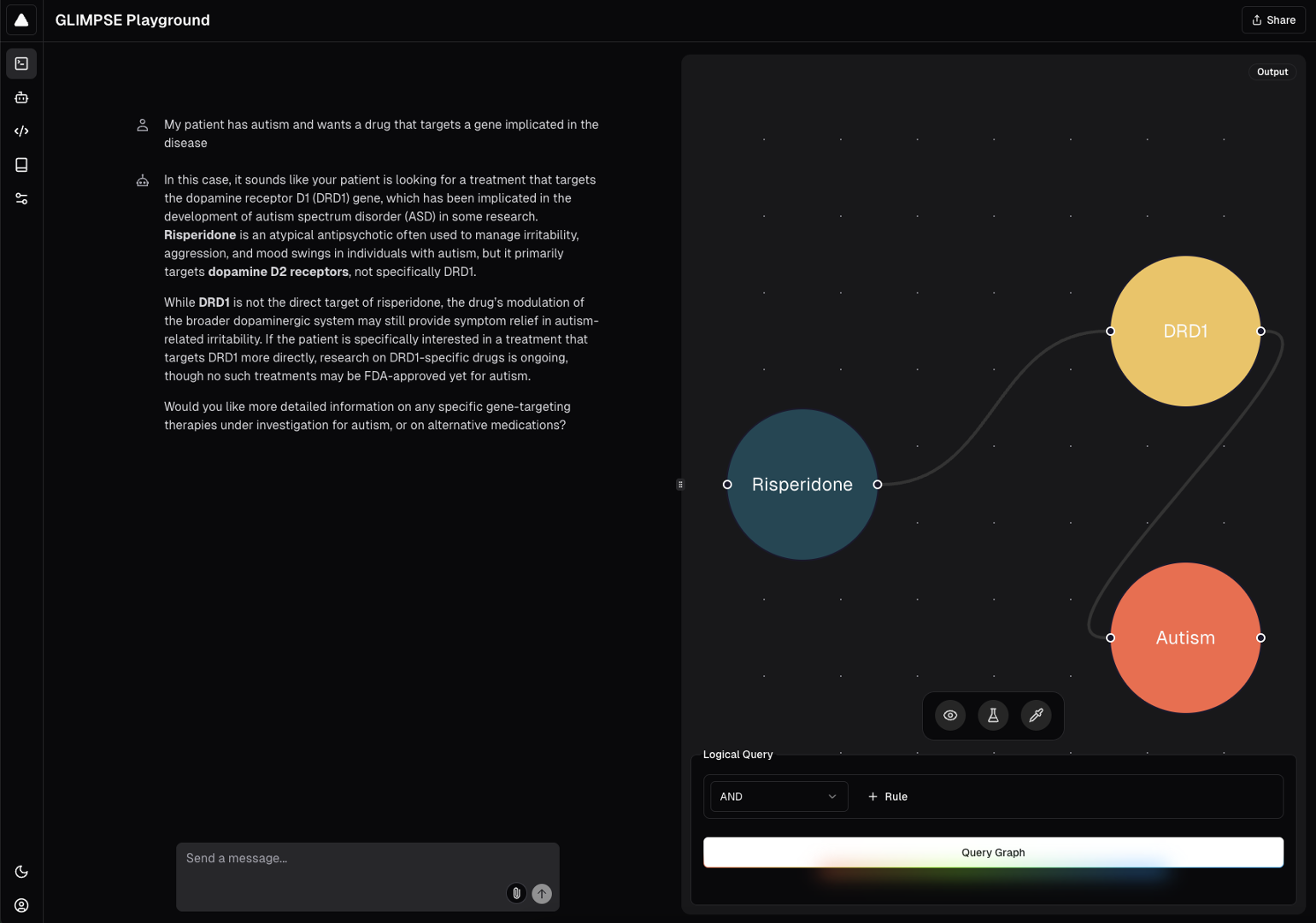
GLIMPSE query with graph visualization.
Background 📚
Link prediction is the problem of predicting the existence of an edge between two entities in a network. Most problems in biology can be formulated as link prediction, e.g., drug repurposing is (drug)—(disease) link prediction; protein network reconstruction is (protein)—(protein) link prediction; drug safety prediction is (drug)—(side effect) link prediction; and so on. Indeed, graph neural networks for link prediction have already demonstrated impressive advances in genetic diagnosis (Alsentzer et al., medRxiv, 2024; Middleton et al., Science, 2024), microbe-drug association (Long et al., Bioinformatics, 2020), drug-drug interaction prediction (Zitnik et al., Bioinformatics, 2018), therapeutic target nomination (Li et al., Nature Methods, 2024), and zero-shot drug repurposing (Huang et al., Nature Medicine, 2024). However, existing methods are only able to predict single edges between two nodes, while clinically and scientifically relevant queries in the real world often require more complex graph queries that involve multiple edges, nodes, and other variables. In the era of precision medicine, complex graph queries would enable contextualized or personalized predictions. For example: "Which combinations of drugs (D_1, D_2, ..., Dn) will simultaneously modulate multiple proteins (P1, P2, ..., Pn) associated with a disease J in a patient with genetic variant V1?”
About GLIMPSE 🤖
To enable complex logical queries on large-scale biomedical networks, we build upon “Embedding Logical Queries on Knowledge Graphs” (Hamilton et al., NeurIPS, 2018), which introduces graph query embeddings (GQE), a method for making predictions about first-order conjunctive logical queries on graph relations. We apply GQE to graph foundation models for biomedicine (e.g., TxGNN, recently published in Nature Medicine) trained on heterogeneous knowledge graphs (e.g., PrimeKG, see Chandak et al., Scientific Data, 2023).
We developed GLIMPSE: Graph-based Logical Inference for Multi-query Prediction and Scientific Exploration, an end-to-end system for complex logical queries on biomedical graphs with applications in contextualized prediction and precision medicine. GLIMPSE is powered by a biomedical knowledge graph with 129,375 nodes and 4,050,249 edges describing biomedical relationships. GLIMPSE includes the following features:
- Smart search across all nodes based on OpenAI text embeddings of node name and type.
- Graph-based retrieval augmented generation to understand and answer general graph queries.
- Smart delegation to a custom GQE-based advanced graph AI model to answer complex precision medicine queries.
- Automatic construction of structured directed acyclic graphs to represent conjunctive logical queries derived from human user-provided free text for GQE prediction.
Implementation Details 💻
We developed a combined front-end and back-end that allows the user to interact with an LLM and with a GQE model through a web interface. Through our interface, a human user can chat with an LLM that has access to the information encoded in the knowledge graph, as well as request additional computation to be performed by external models that specialize in a given biomedical task. The framework that we built is extremely flexible, allowing for any graph and any external models to be added in the future.

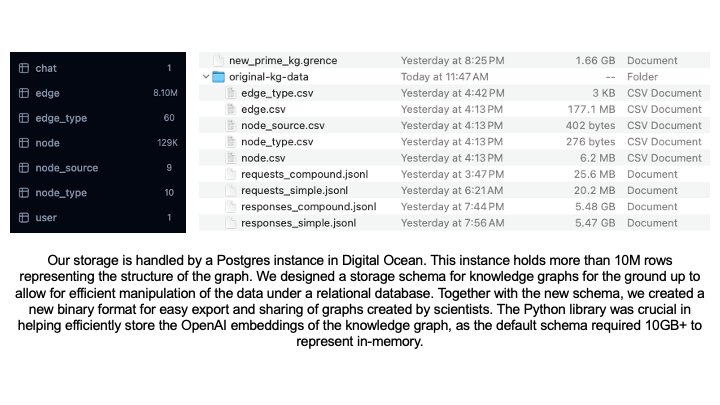
Our storage is handled by a Postgres instance in Digital Ocean. This instance holds more than 10M rows representing the structure of the graph. We designed a storage schema for knowledge graphs for the ground up to allow for efficient manipulation of the data under a relational database. Together with the new schema, we created a new binary format for easy export and sharing of graphs created by scientists. We developed libraries in both Python and TypeScript for loading, saving and exporting knowledge graphs.

We used these libraries internally to develop the rest of the project. The Python library was crucial in helping efficiently store the OpenAI embeddings of the knowledge graph content, as the default schema provided by the company required 10GB+ to represent in-memory. Obtaining the embeddings was done through a batching mechanism with smart retries that sent batched requests to the live OpenAI inference API and monitored the response codes to detect throttling or instances where we were reaching our request quota. It would back down momentarily, slowing down the request rate, and then speed up once no more throttling was detected. This enabled us to generate embeddings over all 129,375 nodes in the knowledge graph in an efficient manner.
The binary representation of the graph was loaded by our backend, written in TypeScript, and uploaded to the SQL database. Due to the memory-intensive nature of the Node.js runtime, loading the entire file at once, even under the new efficient encoding schema, proved impossible. We worked around this constraint by using a two-tiered batching mechanism. The first tier consisted of opening in memory parts of the file in batches, so as to prevent out-of-memory errors. The second tier consisted of subdividing these in-memory batches to send to the database for insertion. All these database operations were managed through the Object Relational Mapper (ORM) Drizzle.js.
We implemented a variety of indexing and search techniques on the database to speed up the analysis on such a large graph. The LLM is able to access information about the nodes in the graph through embedding similarity search (RAG). Humans are able to search through by using full text search over node properties.
Our joint backend-frontend is written in TypeScript using the Next.js framework and React. We rely on the Vercel AI SDK for any embedding and language model-related needs, on Radix UI to create accessible components that make the web interface usable by people of all abilities, and on xyflow to render graphic representations of the knowledge graph. Lastly, we use tailwind and shadcn-ui to create stylish components that adapt to all screen sizes and render correctly in desktops, tablets and phones. Communication between the backend and frontend was accomplished through both traditional HTTP API requests as well as through the use of React Server Components.
The LLM that the user interacts with is also able to call on tools to perform more advanced computation. It relies on the RAG technique described above and vanilla search techniques to find universal identifiers for the entities the user is requesting more information about (e.g., proteins, pathways, diseases, etc.) and constructs a query to the GQE inference service for execution. Once the results are received, they are parsed by the backend and displayed graphically through another pane in the same web interface. Users are therefore able to enter a query entirely in natural language, and this LLM tooling accesses relevant database information to construct the GQE query.
For the specialized graph querying tool, we generated embeddings over the PrimeKG knowledge graph using TxGNN, a therapeutics-centered graph foundation model capable of zero-shot prediction. We aligned these embeddings with the PrimeKG knowledge graph and used them to train a GQE system for multi-node and multi-edge logical query evaluation. We connected the GQE system to the GLIMPSE web interface – this is a work in progress.
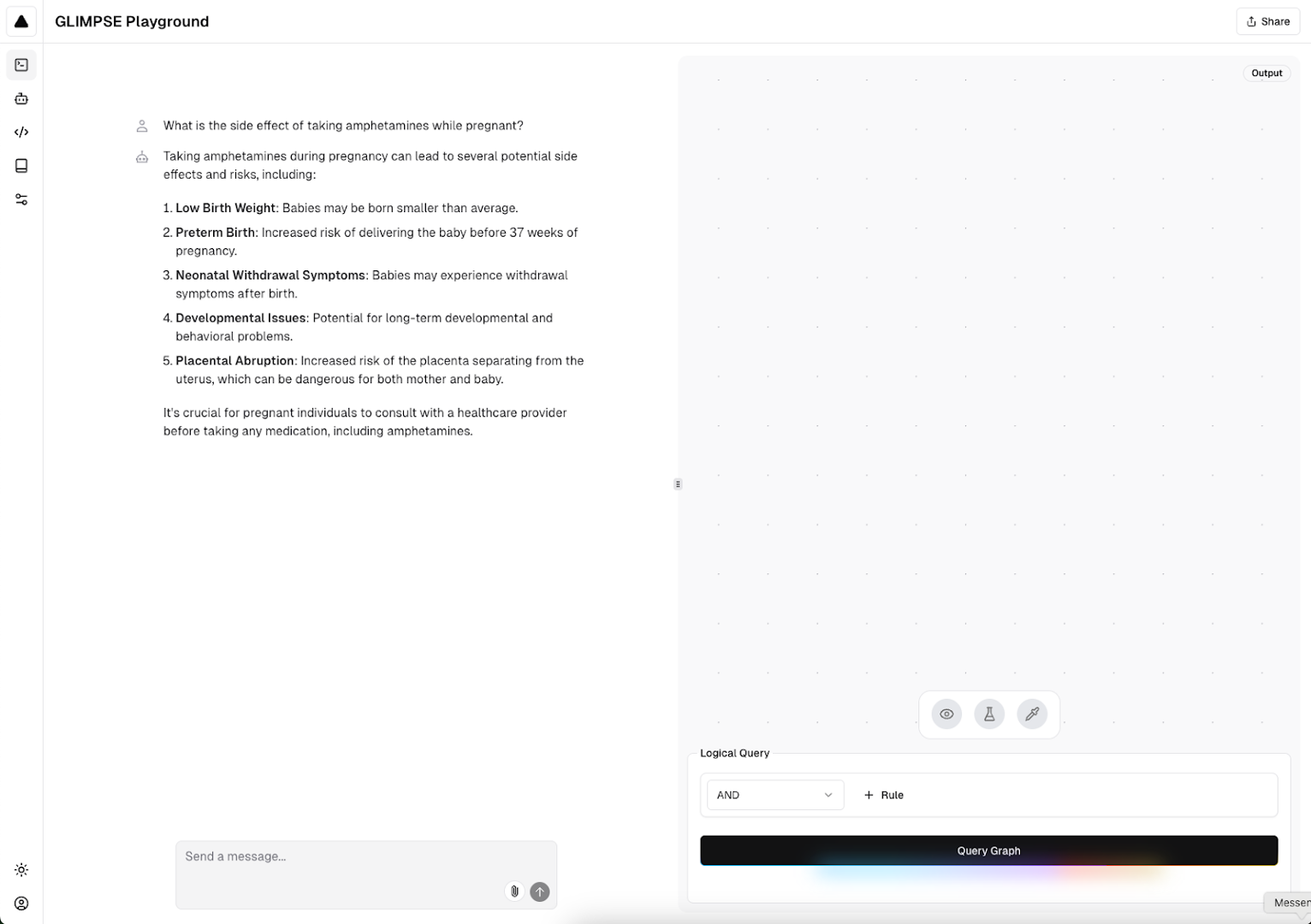
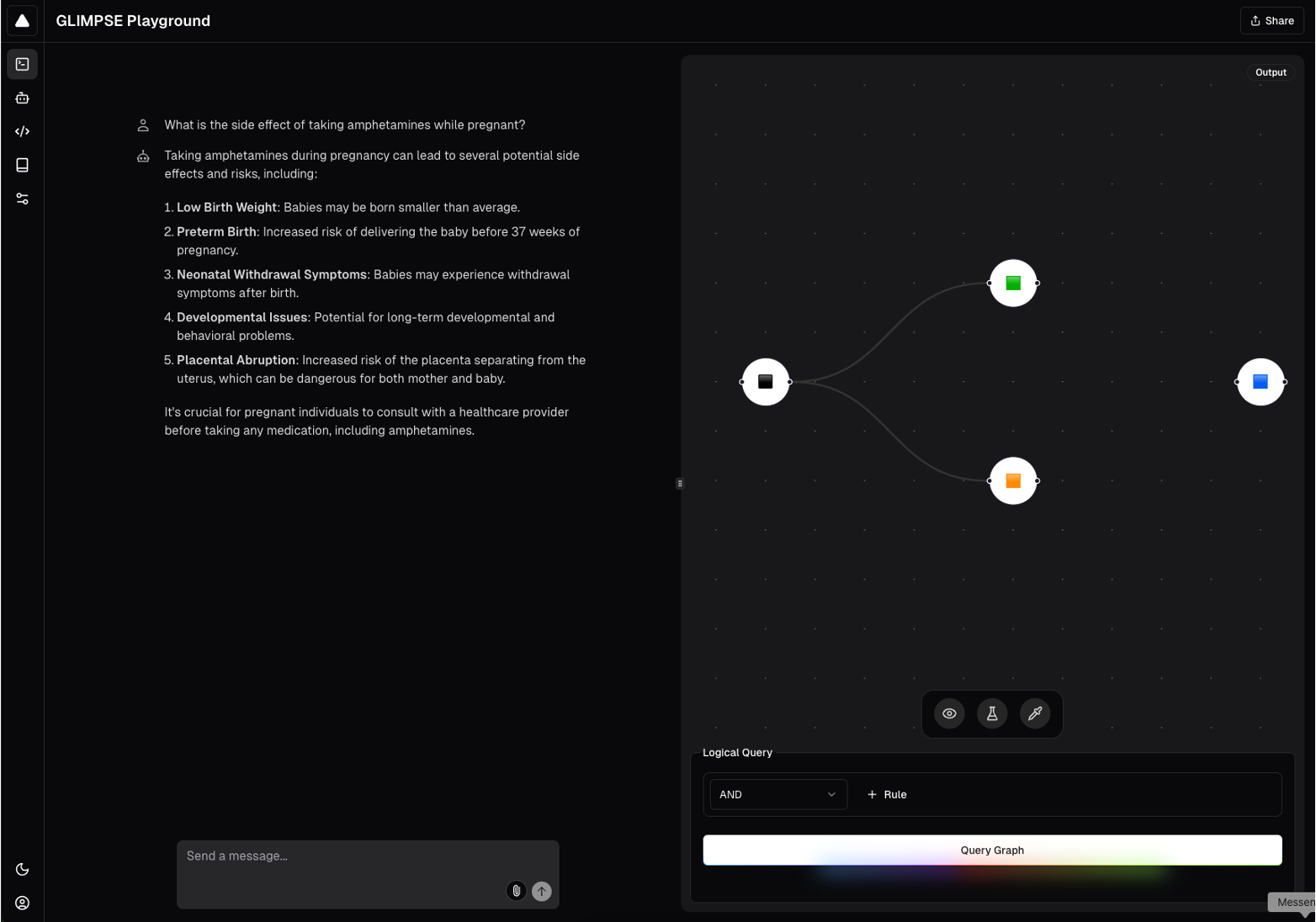
Finally, we developed a pipeline that analyzes graphs in the form of adjacency lists and labeled nodes and relationships and reformats them into three dictionaries: one containing information about edge relations, one with adjacency lists, and one with node type information. We use these three dictionaries to algorithmically sample and generate possible query DAGs from the knowledge graph. We generated over 3 million training queries from PrimeKG, and split the training and test set to ensure that no queries across the training and test sets shared the same target and source node. 70% of the generated DAGs were in the training set, 20% were in the test set, and 10% were in the validation set.
Finally, we evaluated GLIMPSE against the drug safety, protein interaction, and drug repurposing challenges using, for example, the TWOSIDES polypharmacy dataset from TDC. We mapped TWOSIDES nodes onto PrimeKG nodes to predicted edges only in TWOSIDES but not in PrimeKG.
Impact 💊
Our compound AI system makes progress towards AI-driven precision medicine. Example applications of GLIMPSE include:
Genetic and environmental interactions: “Given a combination of genetic mutations (G1, G2, …, Gn) and environmental exposures (E1, E2, …, En), what is the likelihood that a patient will develop asthma?”
Combination therapeutics: "Will a combination of two drugs (D1, D2) target multiple proteins 8(P1, P2)* associated with a genetic mutation (G1) in Alzheimer’s disease?"
Cell type-specific pathology: "Which immune cells (C1, C2) will interact with a viral protein (V1) to trigger an inflammatory response in patients with a specific genetic marker (G1)?"
Universal treatments in patients with comorbidities: “Which drugs can simultaneously target diseases (D1, D2, …, Dn)8 in patients with symptoms *(S1, S2, S3)?”
Overcoming treatment resistance: “For a patient with ovarian cancer associated with specific mutations (G1, G2, G3) and resistance to treatments (T1, T2, …, Tn), what are potential new targets or drug candidates?”
We plan to continue working on GLIMPSE to enable many more complex multi-node and multi-edge queries. GLIMPSE will provide a glimpse into the future of AI-guided personalized medicine.
Development Team ☕
- Ayush Noori (anoori@college.harvard.edu, @ayushnoori)
- Iñaki Arango (inakiarango@college.harvard.edu, @inakineitor)
- Richard Zhu (rzhu@college.harvard.edu, @fiftytwo52)
- Eva Tuecke (evatuecke@college.harvard.edu, @eva.ht)
- Robin Pan (rpan@college.harvard.edu, @redlittlebirdy)
- Vihan Lakshman (vihan@mit.edu, @vihan29)
- Olivia Tang (oliviat@mit.edu, @oliviatang)
- Emmanuel Mhrous (emmanuel.mhrous@princeton.edu, @emmanuel_mhrous)
- Divya Nori (divnor80@mit.edu, @divyanori)
- Andrei Tyrin (atyrin@mit.edu, @andreityrin)
- Bridget Li (bridgetjli37@gmail.com, @bridgetli)
Built With
- css3
- drizzle.js
- html5
- javascript
- node.js
- polars
- postgresql
- python
- pytorch
- radixui
- react
- sql
- tiktoken
- typer.js
- typescript
- vercel







Log in or sign up for Devpost to join the conversation.