Inspiration
I wanted to try a different approach for this hackathon. Rather than find some large available dataset, build a model and train it using the dl1.24xlarge instances to come to the unsurprising conclusion that "Yep, the new Gaudi HPUs are fast", I wanted to address an issue that I noticed with the documentation: The extreme use of ClickOps, which I believe was a bit of a contribution in myself and other peoples having issues on the slack page since the documentation leaves you just enough room to make mistakes if you're not experienced with AWS.
Based on these difficulties I faced, I decided to set my self a challenge of setting up a cloudformation template of the training process of one of the reference models, in the hopes that people can build upon a working cloudformation template for their own training, avoiding them having to same the same problems that I found during the process.
What it does
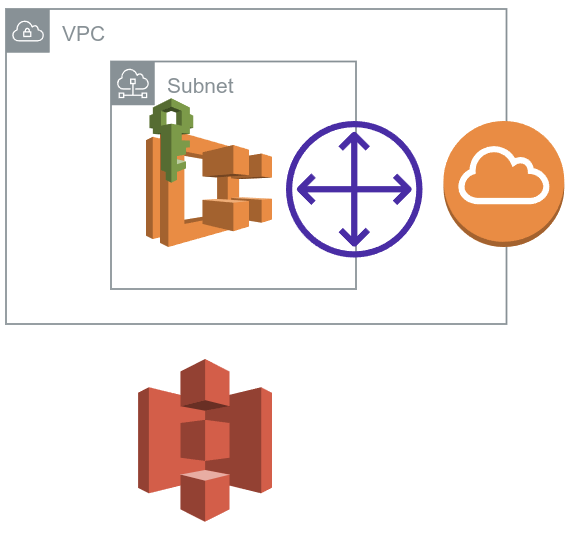
Architecture of cloudformation template.
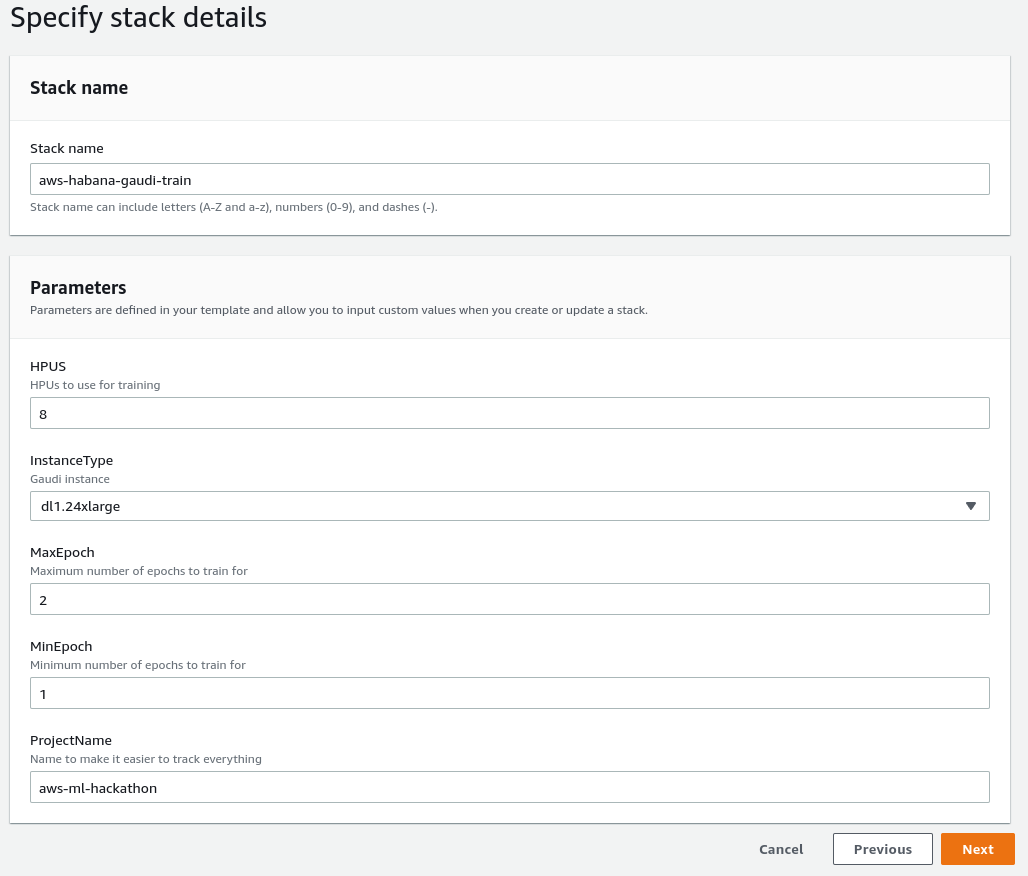
Stack creation screen.
This cloudformation template sets up all the resources required to train the Pytorch Unet and executes a script which trains the model using the Brain Tumor dataset from the Medical Segmentation Decathlon.
As this script is running as UserData progress can be monitored with
tail -f /var/log/cloud-init-output.log
This training script uses all 8 Gaudi HPUs
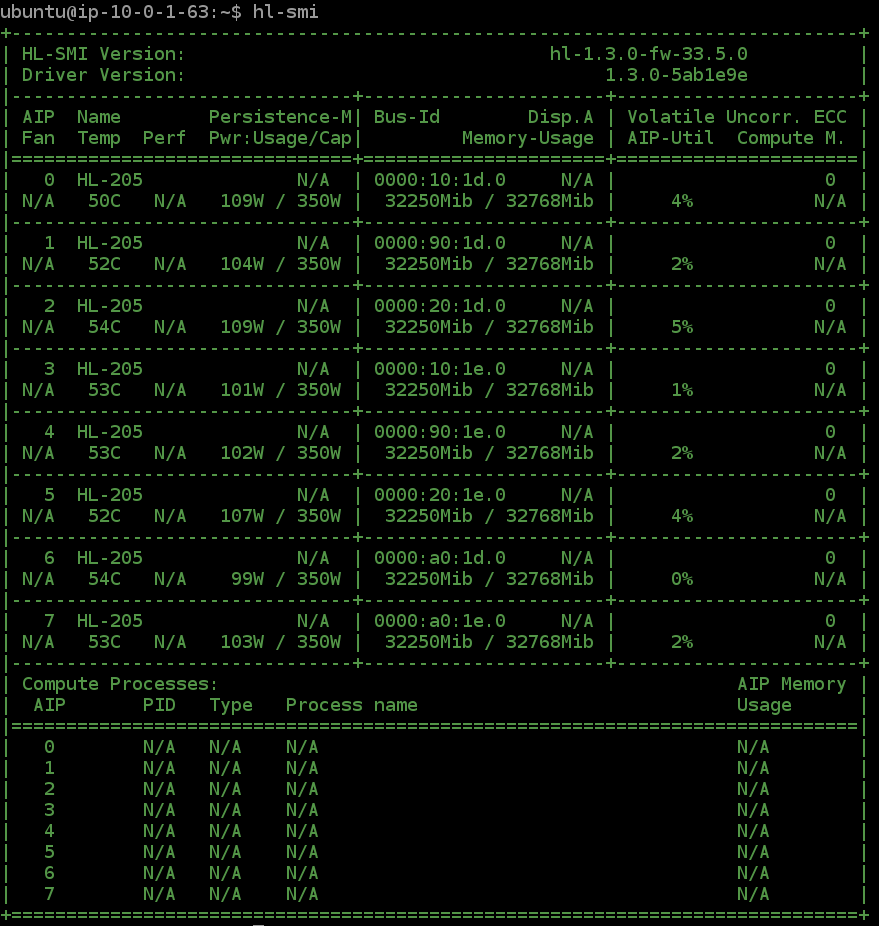
How to use
WARNING: These instances are expensive! Make sure to delete them after you are done!
If you haven't already you will need to raise a service request
for an increase in Running On-Demand DL instances to a value of 96 to run a single dl1.24xlarge instance for the
region that you want to launch this in (Currently only us-east-1 and us-west2 are supported.)
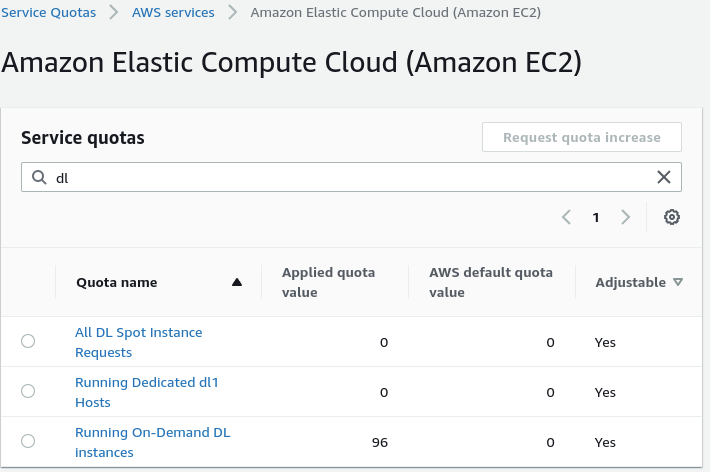
If you cloudformation template gives you the error You have requested more vCPU capacity than your current vCPU limit of 0 allows for the instance bucket that the specified instance type belongs to. then this hasn't been done properly.
Once you're able to request dl1.24xlarge instances, you can then launch the stack either with the buttons below or manually via the AWS CLI.
Launch Stack in US East 1
Launch Stack in US West 2
Or run with the AWS CLI
Or install the AWS CLI,
switch your aws region to aws-east-1 or aws-west-2 and run
aws cloudformation create-stack \
--template-body file://cloudformation/ec2.yml \
--stack-name aws-ml-hack \
--capabilities CAPABILITY_NAMED_IAM
Monitor the stack creation process in the cloudformation page or
monitor the progress in the AWS CLI with aws cloudformation describe-stacks
Once complete visit the EC2 instances page and use Instance Connect to SSH into your instance. In the event that this fails, try again in a few minutes because the instance will show as online before you can access it.
Once there you can view progress of the installation/training process with tail -f /var/log/cloud-init-output.log
The trained model and logs can then be found in /dev/shm/Model-References/PyTorch/computer_vision/segmentation/Unet/output_results
You can then upload these to the S3 bucket that was generated with your cloudformation template since the EC2 instance has an IamInstanceProfile that allows this.
aws s3api put-object --bucket your-bucket-name --key your-file-name
Be sure to empty your bucket with aws s3 rm s3://bucket-name --recursive
if you have added objects to the bucket before deleteting or you'll get a stack deletion failed error. In my experience
your EC2 instance still gets deleted despite this, although you may want to cofirm by visiting instances in the
AWS console for the region you created your bucket in.
Dataset used
This uses the Medical Segmentation Decathlon dataset hosted on AWS OpenData. The dataset is best described in the A large annotaed medical image dataset for the development and evaluation of segmentation algorithms paper.
Specifically though, this training set uses the Task01_BrainTumour dataset which is a set of 750 labelled MRI images from
patients diagnosed with glioblastoma or lower-grade glioma.
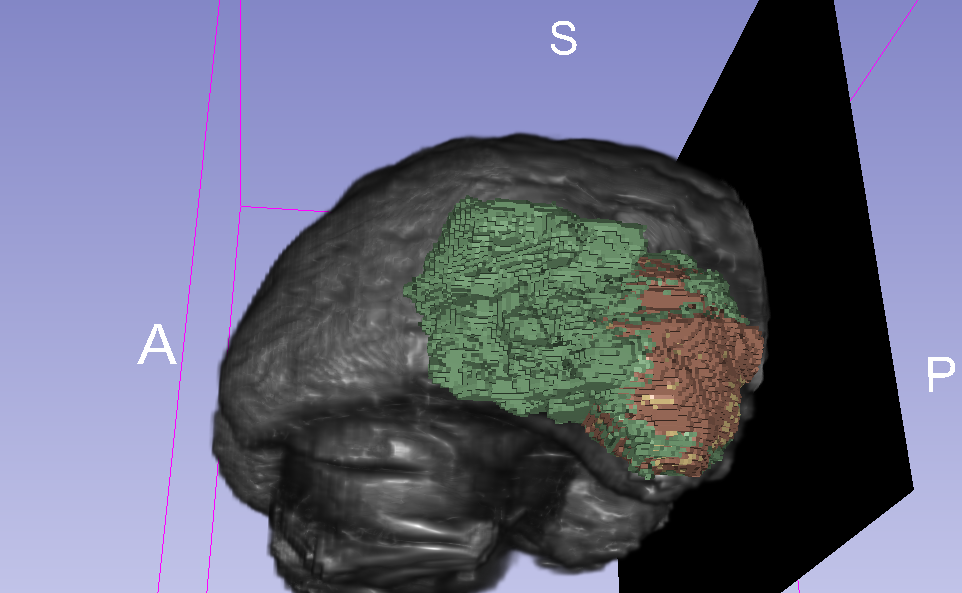
This image was created by loading the BRATS_001.nii data into 3D Slicer.
To speed up training time, I'm running it in 2D mode, which takes each slice independently rather than trying to predict voxel space.
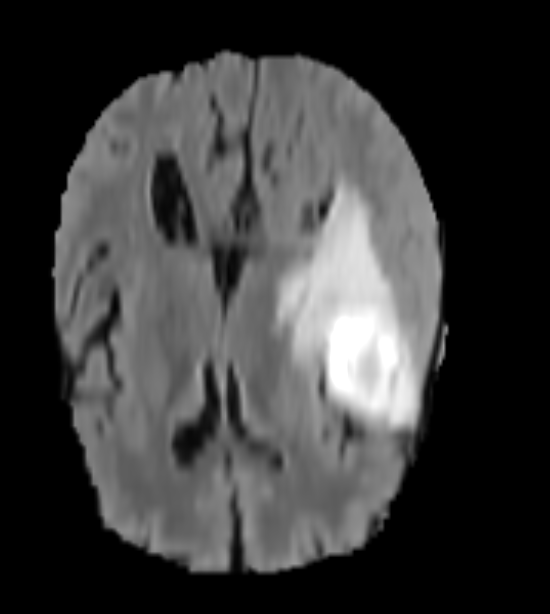
Unlabelled Image
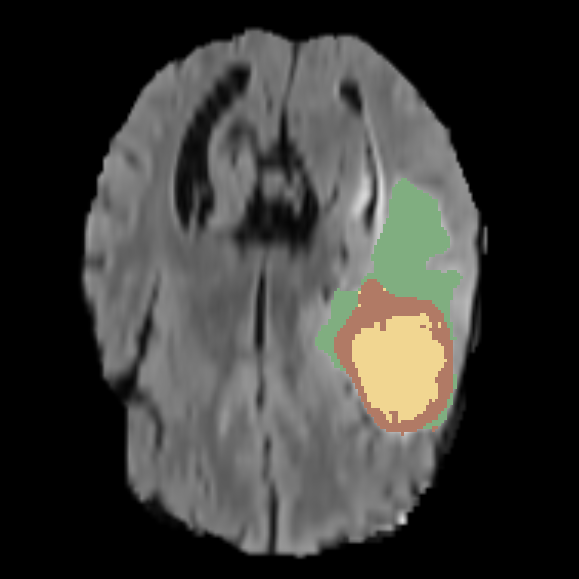
Labeled Image
How I built it
Cloudformation and a LOT of destroying the cloudformation stack and recreating it.
Challenges I ran into
Cloudformation Support
Because the dl1.24xlarge instances are so new, there is limited cloudformation support. This lack of cloudformation has limited me to only trying to get this working via EC2 as the blockers for other services are:
- There is currently no sagemaker support for the new dl1.24xlarge instances
- There is currently no way to enable the correct runtime via ECS in CloudFormation.
### Disk Mounts
Trying to change the root volume in cloudformation when you want to only use UserData is annoying, and the root volume in
the AMI I was using only had
8GBof storage space.
ubuntu@ip-10-0-1-63:~$ df -h
Filesystem Size Used Avail Use% Mounted on
/dev/root 7.7G 3.2G 4.5G 42% /
devtmpfs 374G 0 374G 0% /dev
tmpfs 374G 444M 374G 1% /dev/shm
tmpfs 75G 1.4M 75G 1% /run
tmpfs 5.0M 0 5.0M 0% /run/lock
tmpfs 374G 0 374G 0% /sys/fs/cgroup
/dev/loop0 25M 25M 0 100% /snap/amazon-ssm-agent/4046
/dev/loop2 62M 62M 0 100% /snap/core20/1328
/dev/loop1 56M 56M 0 100% /snap/core18/2284
/dev/loop3 68M 68M 0 100% /snap/lxd/21835
/dev/loop4 44M 44M 0 100% /snap/snapd/14549
/dev/loop5 62M 62M 0 100% /snap/core20/1361
/dev/loop6 44M 44M 0 100% /snap/snapd/14978
/dev/loop7 27M 27M 0 100% /snap/amazon-ssm-agent/5163
/dev/loop8 68M 68M 0 100% /snap/lxd/22526
tmpfs 75G 0 75G 0% /run/user/1000
The main workarounds that I did to bypass this is:
- Pip defaults to
/tmp/for temp files, which when downloading things like pytorch fills up fast. The solution to this was to use theTMPDIRenvironment variable to set pips tmp location to somewhere with more space (/dev/sdm). - The pip cache at
$HOME/.liband the pythonsite-packagesat$HOME/.libcan get a bit big, I symlinked them to another folder mounted in/dev/sdm. - To make sure I don't get any more problems, I just symlinked
openmpiandhabanalabsdirectories since they were large.
UserData and Environment Variables
Turns out UserData doesn't load the variables from /etc/profile.d/habanalab.sh which caused some issues that were VERY hard to track down because everything worked fine when the train command was run manually. You can see in the userdata where I started throwing things at the wall.
I ended up just manually exporting the required variables, it doesn't look the prettiest but it works
Accomplishments that I'm proud of / What we learned
- I was pretty happy that I'd managed to get this working, I haven't done a huge amount of cloudformation before.
- I've never cloudformationed an EC2 instance before, and I became really glad I did when I kept destroying the instance when I started continuously destroying and recreating instances to try to debug issues and check that my fixes worked.
- I learnt a bit about where pip actually stores its files and the files it stores, because I'd never thought about that.
What's next for Quick(er) Start
It'd be worth rewriting some of the code for the userdata into a single makefile so that then you could start selecting different frameworks and different models using just an AWS Cloudformation parameter.
There's some last minute gross things in the UserData that could be cleaned up, you'll see it pretty obviously if you see the exports.
Built With
- amazon-ec2
- amazon-web-services
- cloudformation






Log in or sign up for Devpost to join the conversation.