-
-
DSPDiagnosis Landing Screen
-
DSPDiagnosis Prediction Screen
-

DSPDiagnosis + CrIMR Poster
-
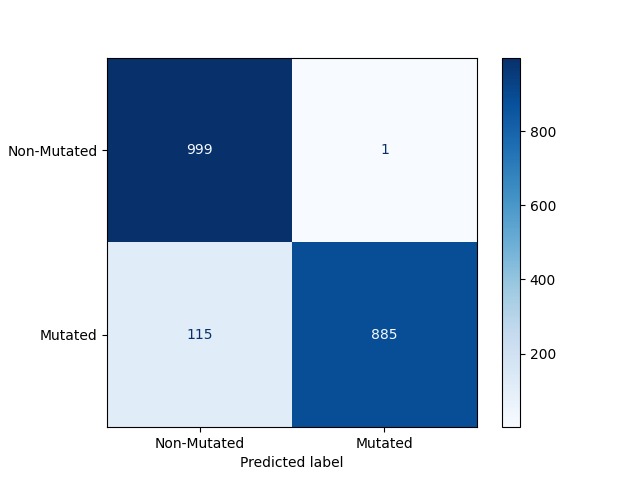
Confusion Matrix for CrIMR
-
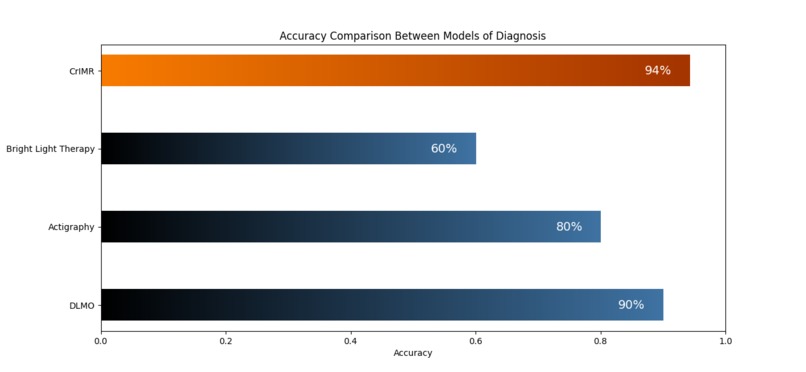
Accuracy Comparison of CrIMR and other models
-
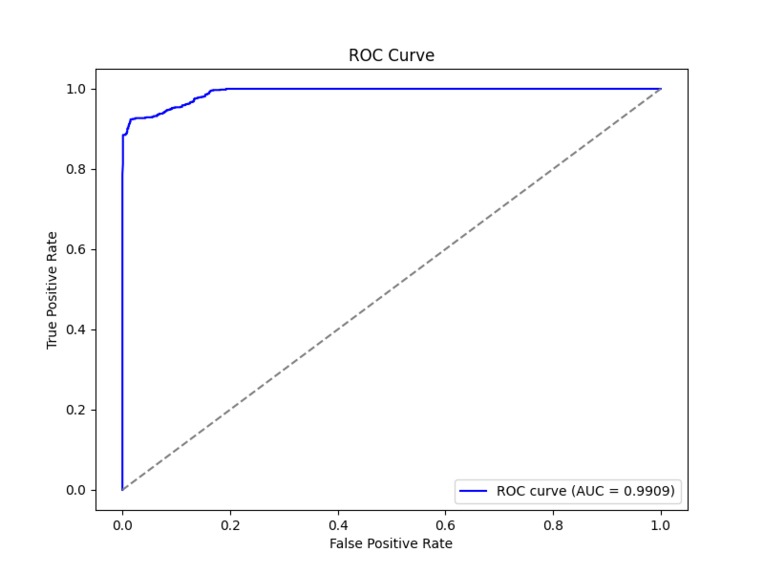
ROC/AUC Curve for CrIMR
-
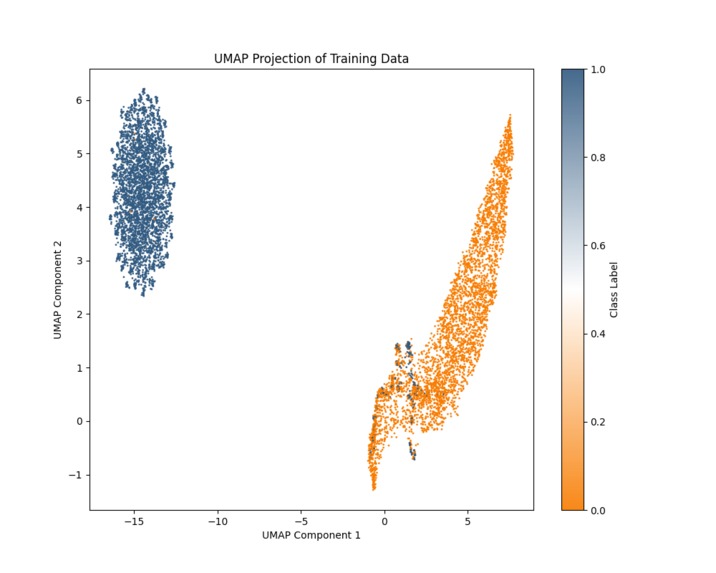
UMAP Dimensional Reduction (trends between mutated and nonmutated are preserved)
-
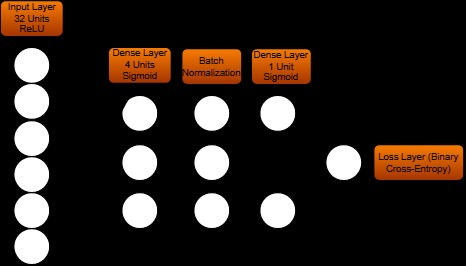
LSTM-based RNN (CrIMR) Structure
-

Parameter Dimensional Reduction
Inspiration
Delayed Sleep Phase Disorder (DSPD) affects up to 16% of adolescents and is linked to mutations in the circadian rhythm, especially in the gene Cryptochrome circadian regulator 1, or CRY1. DSPD is often portrayed as a shift in the sleep-wake cycles, with many facing late sleeping times. I personally was interested in DSPD since I too struggle with sleep and have the symptoms of DSPD, and it is a very common problem among teenagers across the world. The current problem lies within diagnosis. Around 40-50% of DSPD cases are genetically caused, however no current methods for diagnosing genetically-based DSPD exist. Current diagnostic tools like actigraphy and DLMO are indirect, expensive, and time-consuming. I was inspired to explore whether DSPD could be predicted directly from a patient’s CRY1 genetic sequence, allowing for earlier, more accessible diagnosis. My goal was to create a biologically grounded model that uses only genomic data to assess risk for DSPD, integrating machine learning and molecular biology to make a meaningful impact in precision medicine.
What it does
My tool, CrIMR (CRY1 Identification of Mutations and Repression), is a deep learning model that takes a patient’s CRY1 gene sequence in FASTA format and predicts their risk of having DSPD. It uses an LSTM-based recurrent neural network to identify mutation patterns associated with the condition. I also developed a web interface, called DSPDiagnosis, to make the tool accessible in a clinical or research setting. Clinicians or researchers can upload a FASTA file, and the web app instantly returns a DSPD risk score based on the presence of functionally significant mutations.
How I built it
I collected CRY1 gene sequences from NCBI GenBank and ClinVar, focusing on samples with known DSPD-linked mutations and normal variants. Each sequence was one-hot encoded and padded to a uniform length, then input into an LSTM-based neural network trained to classify DSPD likelihood. The model was built using PyTorch, with separate validation and test sets to assess generalization.
To support clinical translation, I built a Flask-based web application that allows users to upload raw FASTA sequences. The server processes the file, feeds it into the trained model, and returns the predicted DSPD risk—making genetic diagnostics more scalable and accessible. The web tool is lightweight, secure, and designed for rapid feedback in a clinical workflow.
Challenges I ran into
The biggest challenge was finding a publicly available dataset. DSPD is one of the less-known disorders and thus, has less datasets centered around the CRY1 gene. However, by using the UCSC genome browser, I was able to sample around 12,000 sequences for training. Another challenge was optimizing the LSTM model to capture long-range dependencies in the genomic sequence without overfitting, which required careful tuning of dropout layers and regularization. Another challenge was the amount of parameters, which exceeded millions and made training memory and time intensive. I solved this using UMAP dimensional reduction, which preserved the trends in the data while decreasing parameters. Finally, integrating a deep learning model into a responsive, production-ready web application required managing PyTorch model serialization and server efficiency for real-time prediction.
Accomplishments that I'm proud of
CrIMR is one of the first machine learning models that enables direct diagnosis of a circadian rhythm disorder from genetic data alone. This project represents a breakthrough in genetics-based diagnosis by eliminating the need for indirect behavioral metrics and offering a scalable, accessible, and personalized diagnostic platform. I am proud that this project earned 1st Place in Computational Biology at the Washington State Science and Engineering Fair (WSSEF) for its innovation in combining deep learning with real-world clinical application.
What I learned
This project taught me how to work at the intersection of machine learning, genomics, and digital health. I learned how to structure DNA sequence data for LSTM input, evaluate model performance in a biological context, and deploy models in a clinical-facing web interface. I also gained experience in full-stack development and API design, which was essential to ensure the tool could be used in research or clinical workflows.
What's next for DSPDDiagnosis: Using the novel RNN CrIMR for DSPD Diagnosis
Next, I plan to expand the system to include other core circadian genes like PER2, CLOCK, and BMAL1, broadening the diagnostic scope to multiple sleep disorders. I also aim to integrate CrIMR into a larger clinical decision support platform that includes genomic analysis, medical history, and phenotype data. Ultimately, my goal is to develop a fast, accessible tool to support genomics-based diagnostics in sleep medicine, particularly for underserved populations where conventional diagnostic testing is limited or unavailable. Furthermore, using this diagnostic model, I aim to integrate functions like the Hill's function and SpliceAI to figure out the impact of the mutation on the fluctuations in repression level of the CRY1 gene to open gateways to personalized gene therapy.











Log in or sign up for Devpost to join the conversation.