What it does
This files takes 49 features related to residue and proteins as input and predict if the residue (row) belongs to a known binding site or not.
How we built it
- used
pythonand thanks to thejupyterhubof Uwaterloo, my computer survives during the training process. - used sklearn and imblearn to process data and train the model
Demo usage - hosted using gradio space
- using pickle to export and import data
host on gradio space
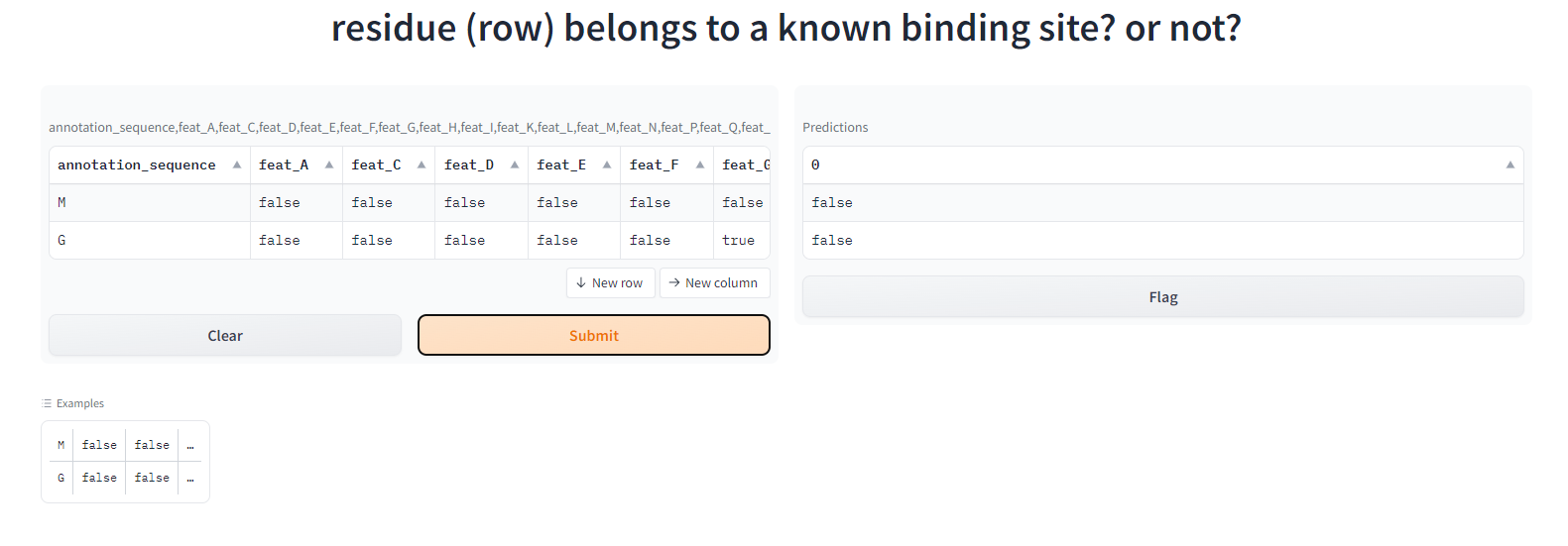
set input parameters
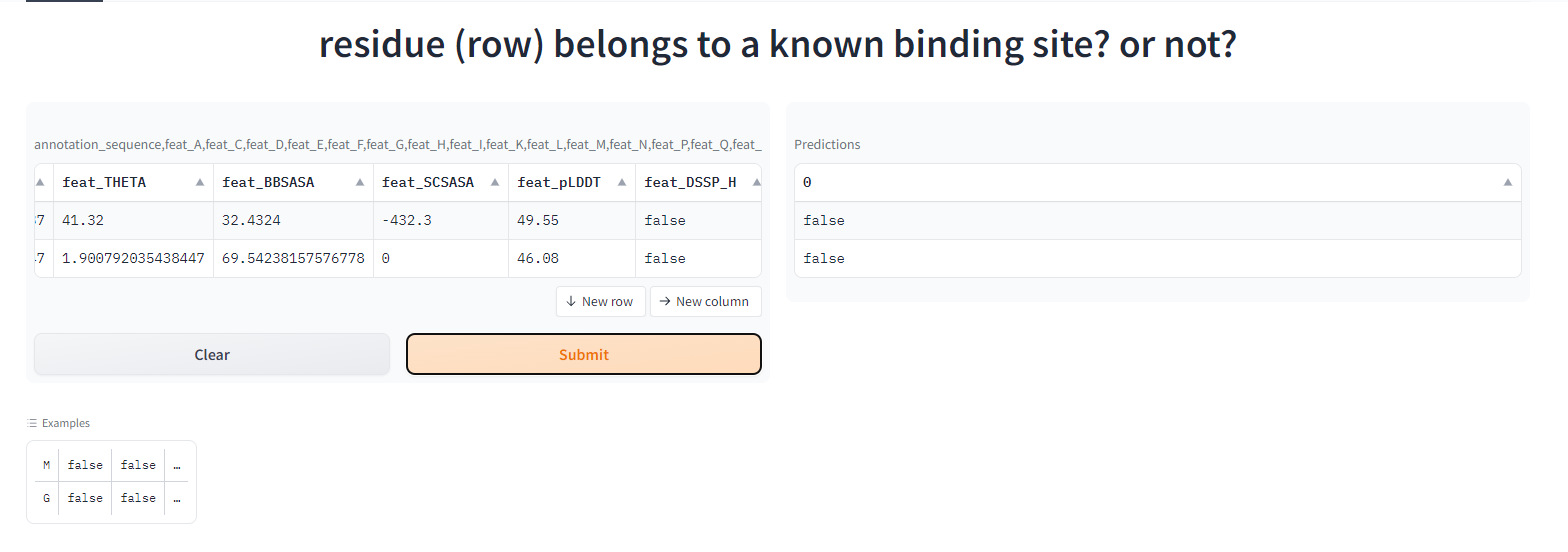
Data Exploration
- checking shape of the dataset (relatively large)
(497166, 50) - notice that the label is binary -> check if imbalanced
False: 479912 True: 17254 basic correlation graph drawn
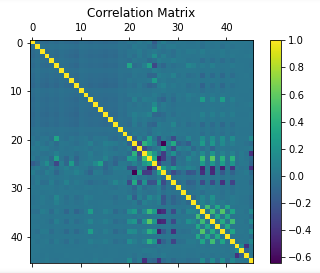

find null values
'annotation_atomrec'
model selection (10 fold 3 repetition cross validaton)
- first under and oversample the dataset by a random amount
- then use some models to compare f1, roc_auc, pr_auc
| model | logistic regression | XGBoost | Random Forest | Neural Network(too slow) | LightGBM | Voting Classifier |
|---|---|---|---|---|---|---|
| f1 | 0.68414 | 0.852539 | 0.8700565 | NA | 0.772295 | 0.860123 |
| roc_auc | 0.85543 | 0.956948 | 0.9664487 | 0.832424 | 0.92824222 | 0.9581390 |
| pr_auc | NA | 0.893542 | 0.90847 | NA | 0.834910 | 0.90321 |
- Random Forest performs the best so we select Random Forest as the prediction model
preprocessing
NA values
- I find column
annotation_sequenceandannotation_atomrecare exactly the same besides thatannotation_atomrechas na values. => dropannotation_atomreccol
scaling
- check numerical data, most of them are not norm distributied => using
minMaxScalar()
encoding categorical data
- don't use one hot encoding because the categories for the categorical varaible is too much => use
LabelEncoder()
check outliers
- bool covaraibles is too heavy on false values (no action performed)
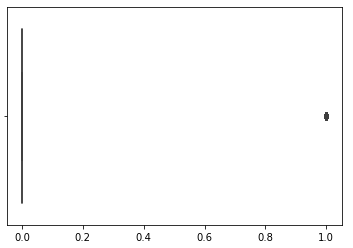
deal with imbalanced data
performed
SMOTEandRandomUnderSampletotunning considering run time efficiency, overfitting and roc_auc pr_auc [a, b] a is the sampling strategy for SMOTE and b is the sampling strategy for RandomUndersample
| sample_strategy | 0.1, 0.5 | 0.1, 0.3 | 0.2, 0.5 | 0.3, 0.6 | 0.5, 0.8 | 0.6, 0.9 | 0.8, 0.9 |
|---|---|---|---|---|---|---|---|
| roc_auc | 0.96861 | 0.972335 | 0.9895 | 0.99462 | 0.9975907 | 0.998134 | 0.99893 |
| pr_auc | 0.90295 | 0.87756 | 0.94928 | 0.968687 | 0.9830687 | 0.98615 | 0.9900327 |
- To avoid too much samples and runtime too slow, choose
[0.5, 0.8]Counter({False: 299945, True: 239956})
model tunning
imblanced = {false:1, true: 1.2}
| parameters | class_weight=imbalanced, bootstrap=False | class_weight="balanced", bootstrap=False | class_weight=imblanced, bootstrap=False, max_features="log2" | class_weight=imbalanced, bootstrap=False, n_estimators=50 | class_weight=imbalanced, bootstrap=False, n_estimators=80, max_features="log2" | class_weight=imblanced, bootstrap=False, min_samples_split=4, max_features="log2" | class_weight=imblanced, bootstrap=False, min_samples_split=3, max_features="log2" | class_weight=imblanced, bootstrap=False, min_samples_split=3, max_features="log2", max_depth=20 | class_weight=imblanced, bootstrap=False, min_samples_split=3, max_features="log2", min_samples_leaf=5 |
|---|---|---|---|---|---|---|---|---|---|
| roc_auc | 0.9958320 | 0.995803 | 0.995837 | 0.995396 | 0.99573 | 0.99576 | 0.995837 | 0.99258643 | 0.995076 |
| pr_auc | 0.985214 | 0.985180 | 0.985201 | 0.98509 | 0.985180 | 0.98520 | 0.98531 | 0.9768451 | 0.984155 |
| f1 | NA | NA | NA | NA | NA | NA | NA | 0.965574 | 0.97235 |
- final choice
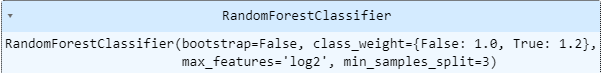
feature importance
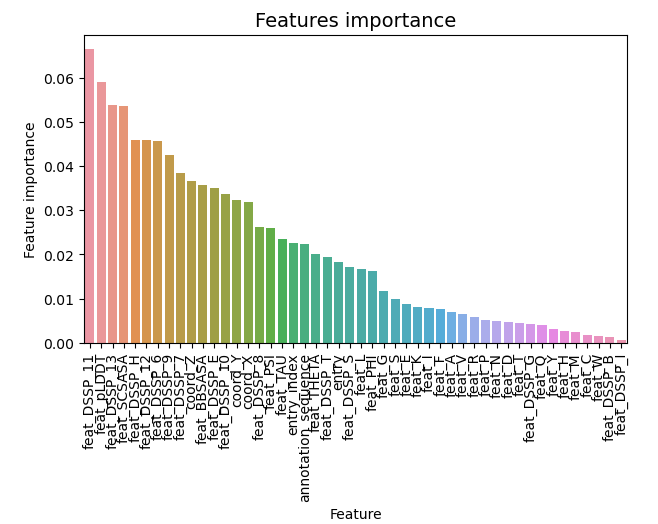
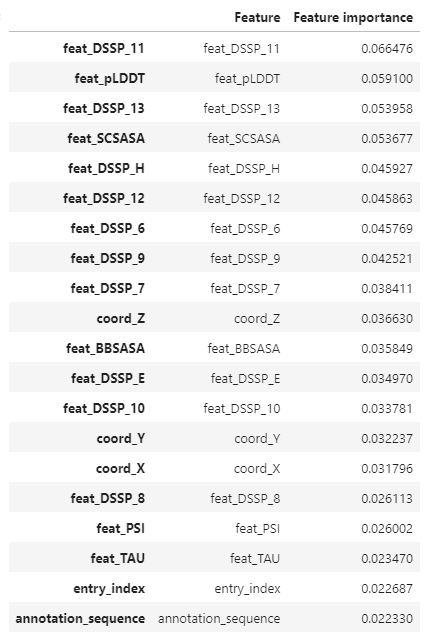
predictions
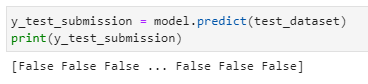
Challenges we ran into
- how to set up gradio space
- the runtime for the model is too long and my laptop died while running it
Accomplishments that we're proud of
- I successfully finish the model with a relatively high accuracy
- I managed to host the gradio space
What we learned
- I have also learned the process for preprocessing data and how to tunning the hyperparameters
- i have learned how to deploy model
Built With
- huggingface
- jupytornotebook
- python
- randomforest
- sklearn




Log in or sign up for Devpost to join the conversation.