Inspiration
As per the World Heart Report 2025, Cardiovascular disease (CVD) remains the leading cause of death globally, with a projected 20.5 million deaths in 2025, rising to 35.6 million by 2050. And the cardiovascular treatment experiences some of the highest rates of adverse drug reactions (ADRs) primarily due to polypharmacy the fact that heart patients typically require multiple simultaneous medications and according to recent pharmacological studies indicating that over 53% of these ADRs are potentially avoidable. ML models trained on massive amounts of therapeutically relevant data may help physicians make more informed clinical decisions before prescribing drugs.
Existing DDI prediction systems rely on manually curated databases covering limited drugs and require manual mapping of new drugs, and memorise interactions instead of learning molecular patterns. I present AushadhiNet-GATv2 , a graph neural network architecture that learns molecular-level interaction patterns from chemical structures
What it does
AushadhiNet-GATv2 is a graph neural network model that predicts drug-pair interactions with probability scores and classifies interaction types (potential adverse drug reactions) across 86 mechanisms. The system alerts physicians and patients before prescribing or taking medications, preventing harmful drug combinations.
I integrate it with streamlit application with manual drugs entry automated OCR-powered image scanning and patient profiling for production-level use and serving end users.
Key features:
- High Accuracy: 90% accuracy and 97.94% recall on 19,000 DDI pair
- Prescription Scanner: OCR-powered image scanning with fuzzy drug name matching
- Risk Profiling: Patient-specific cardiovascular risk stratification validated on 70,000 CVD patient records
- Real-time Inference: <100ms prediction latency, deployable on consumer hardware
- Multi-drug Analysis: Checks all pairwise combinations for upto 4 concurrent medications
- Extensible Architecture: Supports fine-tuning on additional datasets (new drugs, rare interactions, institution-specific data) for continuous accuracy improvement
How I built it
Data Pipeline:
I constructed a comprehensive drug interaction graph from DrugBank v5.1.9, containing 1,706 medications, molecular structure and 191,808 documented interactions. For each drug, we extracted three complementary molecular representations:
- Morgan fingerprints (1024-dim) capturing local substructural patterns
- MACCS keys (167-dim) encoding functional groups
- Physicochemical descriptors (8-dim) quantifying global molecular properties
Additional integration with hackathon official cardio_bas.csv data during inference.
Model Architecture:
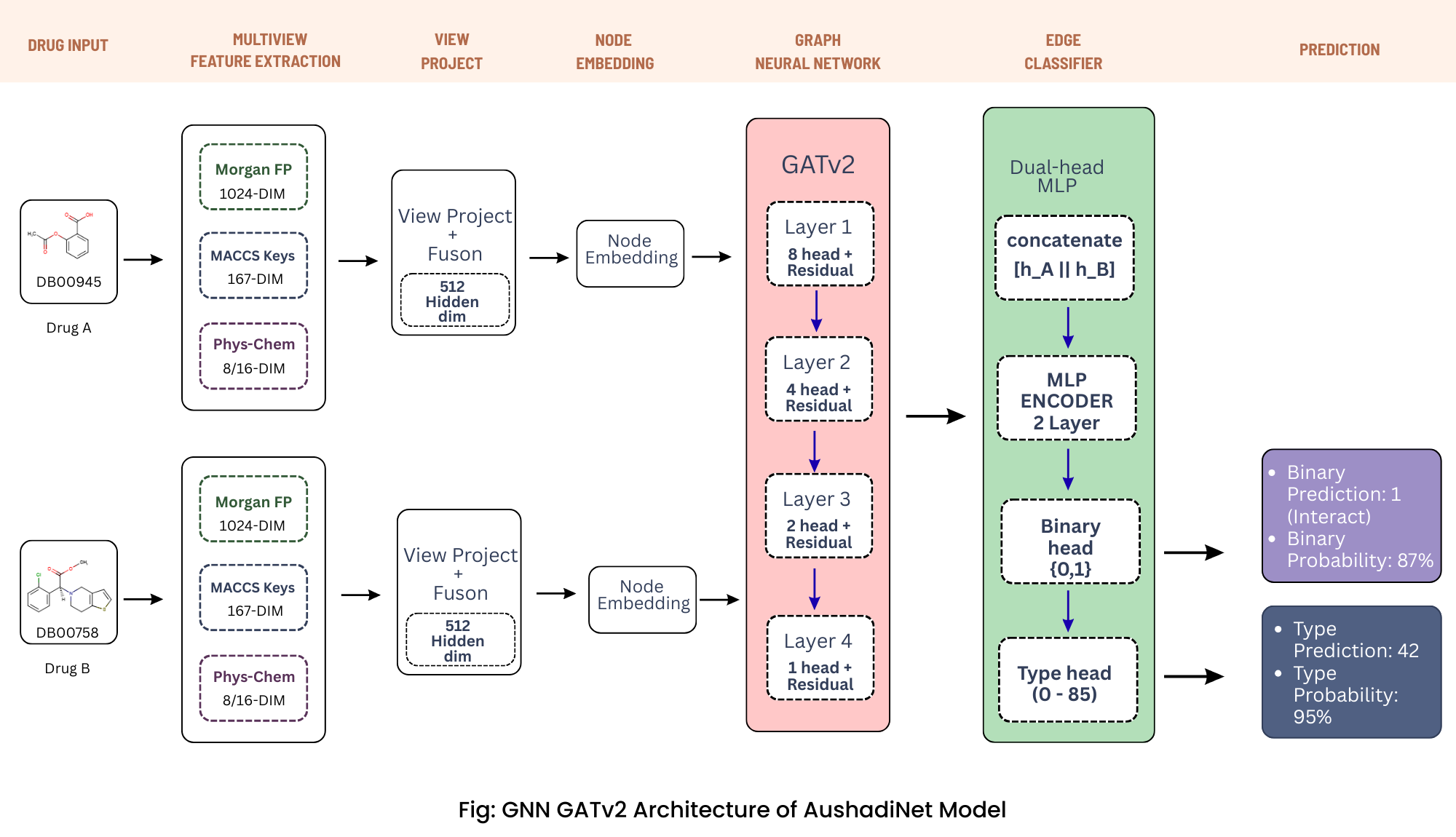
AushadhiNet-GATv2 employs a latest and advance multi-view learning framework:
- View Projection: Three parallel MLPs transform heterogeneous molecular features into a unified 512-dimensional space
- Attention Fusion: Learned attention mechanism weights the importance of each molecular view
- Graph Convolution: Four stacked GATv2 layers with residual connections perform hierarchical message passing
- Dual-Head Classification: Predicts both interaction probability (binary) and mechanism type (86 classes)
Training Strategy:
- Dual-optimizer approach with gradient accumulation (effective batch size: 2048)
- Focal loss to address class imbalance or customizable loss
- Data augmentation via edge dropout and feature perturbation
- Active model tracking system to save the best model
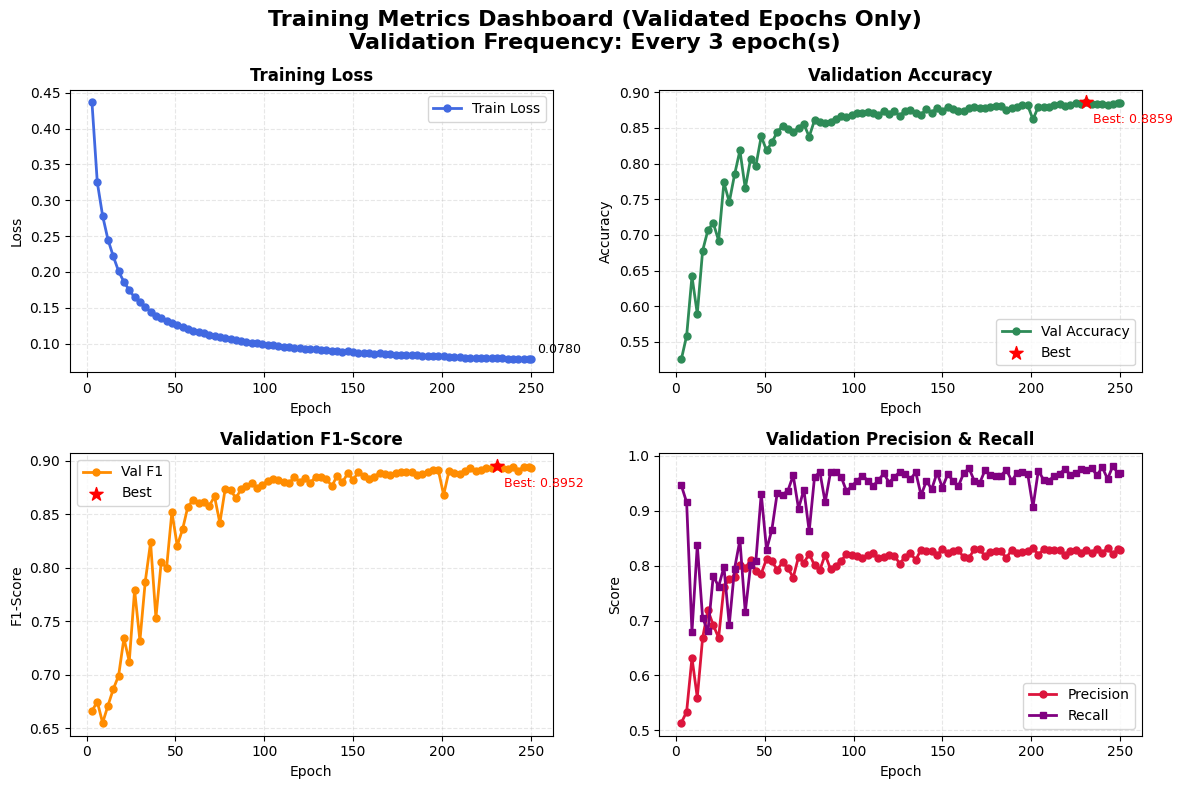
- Comprehensive four-panel metrics dashboard tracking key performance indicators across validated epochs
Deployment:
Built a Streamlit web application to immediately serve the end user with features:
- Manual drugs entry
- OCR integration for automated prescription image scanning and prediction
- Fuzzy drug name matching
- Patient risk profiling using age, blood pressure, cholesterol, and smoking status
- Real-time molecular structure visualisation with RDKit
- Clinical drug information fetched from PubChem API
- Mapping the interaction type to provide the ADR description
Challenges I ran into
- Accuracy Plateaus and Architectural Limits: Experimented with multiple GNN architectures (GAT, GCN, GraphSAGE) over 1.5 weeks to break through performance limits, and achieved maximum accuracy with faster training with GATv2's Dynamic Attention mechanism.
- Computational constraints: I frequently ran into
Out of MemoryandGoogle Colab GPU Usage Limits, requiring optimisation strategies like gradient accumulation, mixed-precision training (FP16), and memory-efficient neighbour sampling. I was constantly switching between Kaggle and Colab for continued training and experiments - Data Quality & Validation: Mapped and validated 86 interaction type descriptions using DrugBank documentation and Medscape clinical references to ensure medical accuracy
- Class Imbalance: Addressed severe imbalance between positive (interactions) and negative (safe). I solved it using focal loss and data augmentation
Accomplishments that I'm proud of
- My first research on the molecular level
- State-of-the-art Performance: Achieved 90% accuracy and 97.94% recall, 92.07% ROC-AUC, outperforming many big-budget funded DDI prediction systems.
- Clinical Validation: Validated risk profiling system against 70,000 real CVD patient records
cardio_base, demonstrating practical clinical applicability - Research Contribution: Documented entire methodology in research paper with mathematical formulations and ablation studies as an independent sole researcher.
What I learned
- Cardiovascular Epidemiology: Understanding of polypharmacy challenges in CVD treatment, where 68% of patients have multiple comorbidities requiring 5-10 concurrent medications
- Graph Neural Networks: Learnt GATv2 architecture, multi-head attention mechanisms, and message passing for molecular interaction learning
- Pharmacology: Understood drug interaction mechanisms, ADME properties, and clinical significance of different DDI types
- I also read about 12 research papers on CVD treatments and polypharmacy, graph neural network and updated GATv2's "dynamic" attention mechanism
What's next for AushadhiNet-GATv2
- Dataset Expansion: Official access to DrugBank latest (2025-26) dataset for fine-tuning on the latest drug interaction data and rare ADR cases
- Enhanced Clinical Intelligence: Develop a sophisticated ADR description pipeline with severity scoring, alternative medication suggestions, and contraindication warnings
- Advanced OCR: Implement transformer-based handwriting recognition (TrOCR) to handle doctors' fuzzy prescriptions and low-quality images
- Brand Name Support: Expand drug database to accept commercial/brand names.
The research report document provides in-depth information.
Built With
- drugbank-dataset
- ocr
- pubchem
- python
- rdkit
- streamlit
- torch







Log in or sign up for Devpost to join the conversation.