-
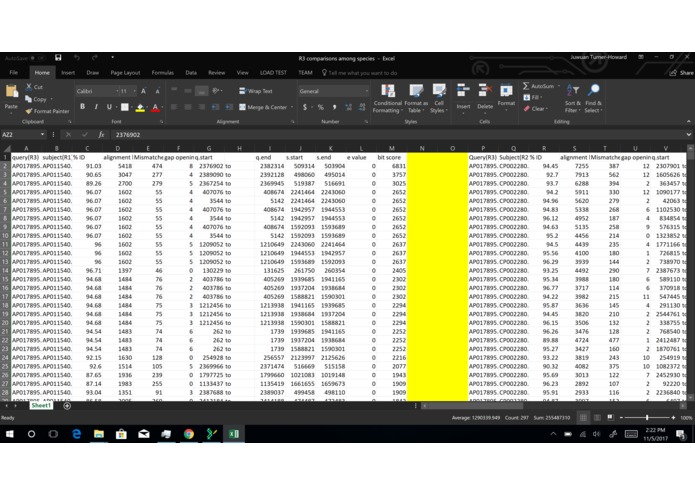
Part 1 on data
-
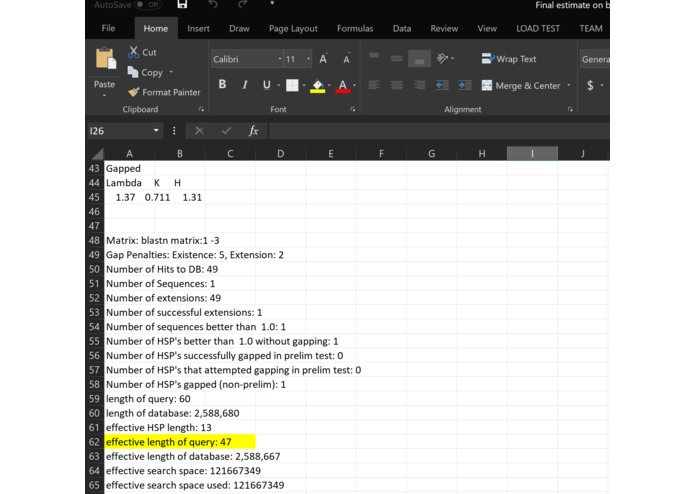
Part 2 on data
-
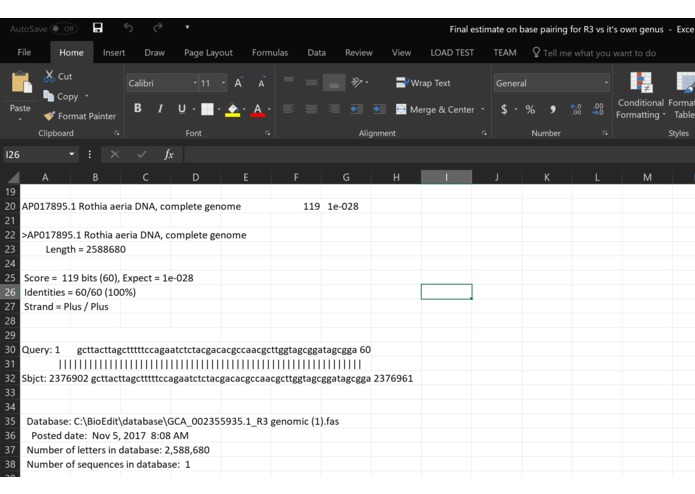
Part 3 on data
Inspiration:
What it does: We have model program, but the program does not work because of lack of data. However, we have a conceptual idea of what it would do if it had enough data. In the beginning, we were all lost, but common knowledge about genetics and genomic analyze we were able to overcome this confusion. Essentially the goal of the program is to take alignments jumbled with the variants from different genomes and identify the individual species based on these variants. From there it would be able to tell not only the classification of the species but also the contamination levels form in the human DNA sample.
How we built it: We built a model program in python, but due to the lack of data we could not properly run it.
Challenges we ran into: sequencing the data and analyzing it completely.
Accomplishments that we're proud of: We pinpointed variants that make a species unique to its original genome locations.
What we learned: That sequencing multiple genomes in an efficient way is very difficult as well as finding a program to help us analyze this data without crashing
What's next for C4Blast: We want to connect with other teams about this challenge and try to come up with a program and a definite answer in tackling this project in its entirety. We want to be able to sequence all the data, not just a part.
Log in or sign up for Devpost to join the conversation.