-
-
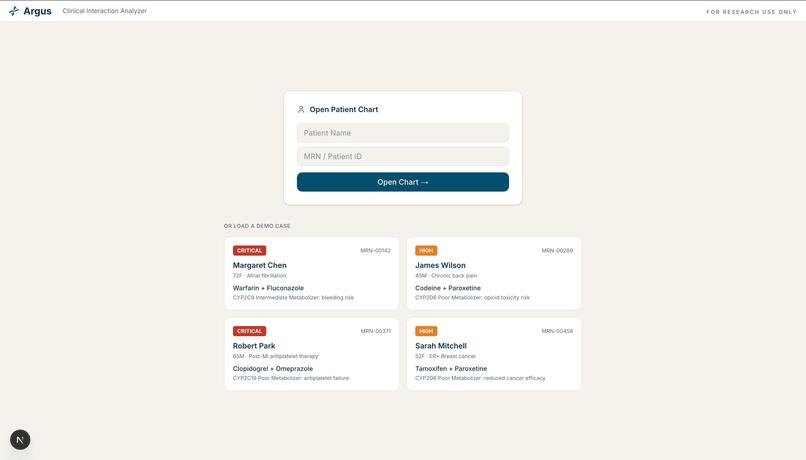
Patient lookup with four preloaded demo cases, each flagged by risk level and drug-gene collision.
-
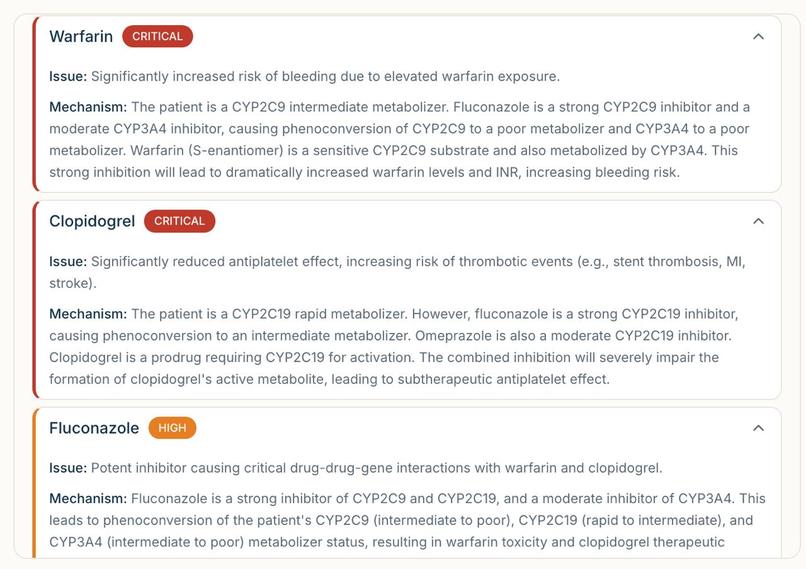
Each drug gets a risk card with its issue and biological mechanism, flagged Critical or High.
-
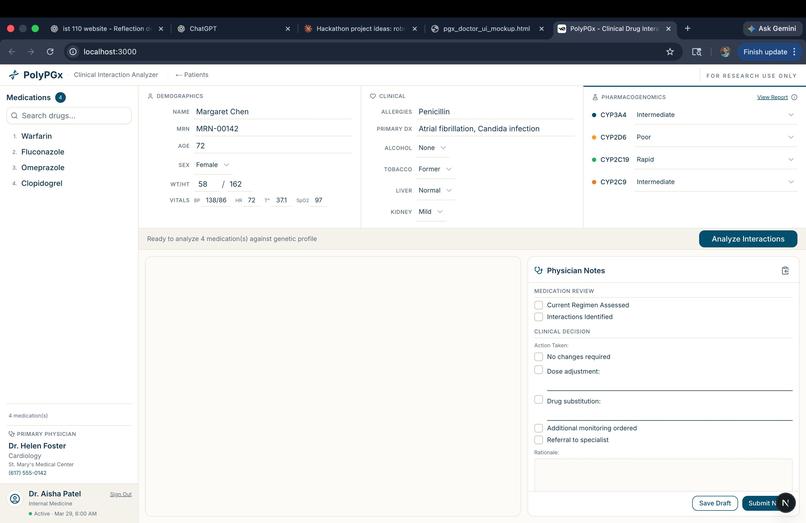
Patient loaded, genetic profile entered, ready to run the full interaction analysis pipeline.
-
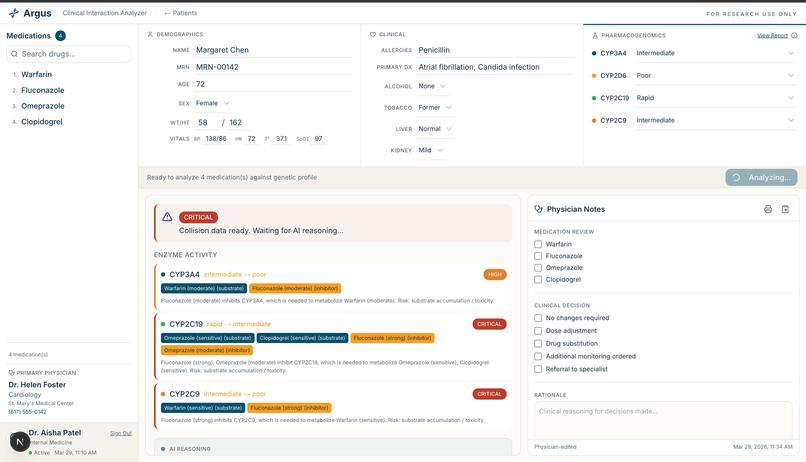
Full patient chart showing real-time enzyme activity, collision detection, and physician notes panel.
"Every time I went to the doctor, they gave me another pill." Argie, a widowed former teacher, started to feel sickly in her mid-70s. Simple mental tasks seemed to exhaust her. She was often nauseous, fatigued, and restless. Still, a memory specialist concluded that her cognitive abilities were intact. The real culprit? The twenty-one vials of medication sitting atop her bedside table.
Argie is not an outlier. Polypharmacy — the simultaneous use of five or more medications — contributes to an estimated 150,000 premature deaths in the U.S. annually. Adverse drug events (ADEs) are now estimated to be the third leading cause of death in the nation (American Society of Pharmacovigilance). ADEs cost the US healthcare system $30 billion a year, with more than 1.5 million people visiting emergency departments. And the risk builds fast: with two medications, the chance of an adverse reaction is around 13%. With seven or more, it surges to 82%. The tragedy is that most of these interactions are biochemically predictable — if you know where to look.
What inspired us to begin this project is a gap hiding in plain sight. Pharmacogenomics (PGx) — the science of how your genes affect how you process drugs — has produced genuinely powerful tools. But almost every clinical PGx tool analyzes only one drug at a time. A patient gets a flag: CYP2D6 poor metabolizer → avoid codeine. Simple. Useful. But incomplete.
Real patients, especially elderly ones, are often on 8–12 medications concurrently. Many of those drugs compete for the same metabolic enzymes —for example, CYP2D6, CYP2C19 or CYP2C9. When a drug inhibits an enzyme that another drug depends on, and the patient's genome already makes that enzyme slow, the effects cascade in ways no single-drug lookup table can capture. That is not a lookup problem. That is a reasoning problem. We built the tool that reasons.
Argus is built on a core architectural insight: logic first, AI reasoning second. The foundation is a CYP Collision Detector. We built a curated database mapping commonly prescribed drugs to their CYP enzyme interactions, sourced from the FDA CYP450 drug interaction table, and validated against PharmGKB. When a clinician enters a patient's medication list, this layer maps every drug to its enzymes, finds overlaps, and produces a precise collision map. It cannot hallucinate. The same input always produces the same output.
Only then does the AI enter. Rather than asking the model to answer from memory, we retrieve the most relevant pharmacogenomics evidence from PharmGKB — the world's most comprehensive pharmacogenomics database — and pass it to the reasoning engine alongside the collision map and the patient's genetic profile. The model is instructed to reason across all drug interactions simultaneously, not one at a time, producing a structured clinical note with risk level, bottleneck enzyme, problematic drug combinations, and recommendations grounded in CPIC guidelines.
The interface is built with Next.js and deployed on Vercel, giving judges a live URL to interact with directly.
Building this taught us that the hardest problems in applied AI are rarely about the model — they are about what you feed it. Prompt engineering the clinical reasoning layer was the most iterative part of the build. Getting the model to reason across a full drug list rather than about each drug in sequence required careful design and biological validation at every step.
We also learned that domain expertise on a team changes everything. Having a biology team member who could check the model's output meant we were building something clinically defensible, not just functional.
Data quality was a major challenge. Manually curating and validating drugs against the FDA CYP450 table and cross-referencing with PharmGKB required real biological expertise and time that a 24-hour hackathon does not generously offer. Getting it right was non-negotiable.
Argie got better, eventually — after a clinical pharmacy specialist worked through her full medication list and brought her down from twenty-one drugs to eight. She was less confused, less fatigued, steadier on her feet. That intervention took a specialist and weeks of time. We built Argus to make that reasoning available in seconds.
Built With
- antigravity
- claude
- claudecode
- fda.gov
- gemini
- ncbi
- pharmgkb
- v0.dev
- windsurf




Log in or sign up for Devpost to join the conversation.