-
-
We ideated and built 10+ biomedical prototypes—from digital cell twins to abstract research interfaces
-
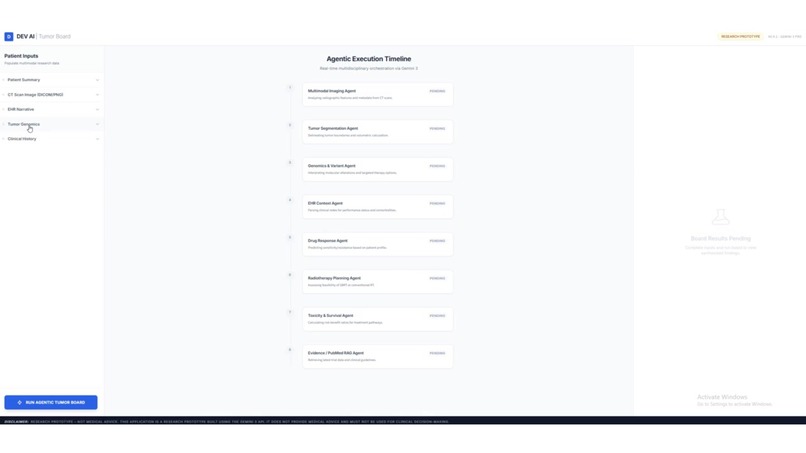
Ideated a virtual tumour board interface for agentic, multi-specialist collaboration.
-
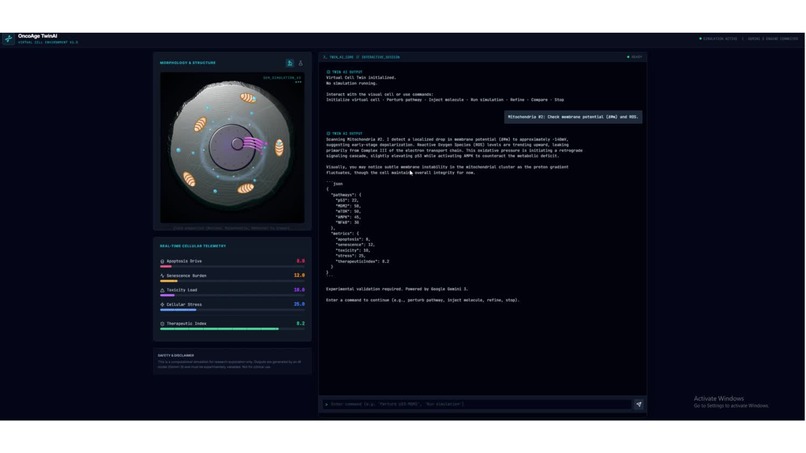
Ideated an interactive virtual digital cell twin for aging and cancer research.
-
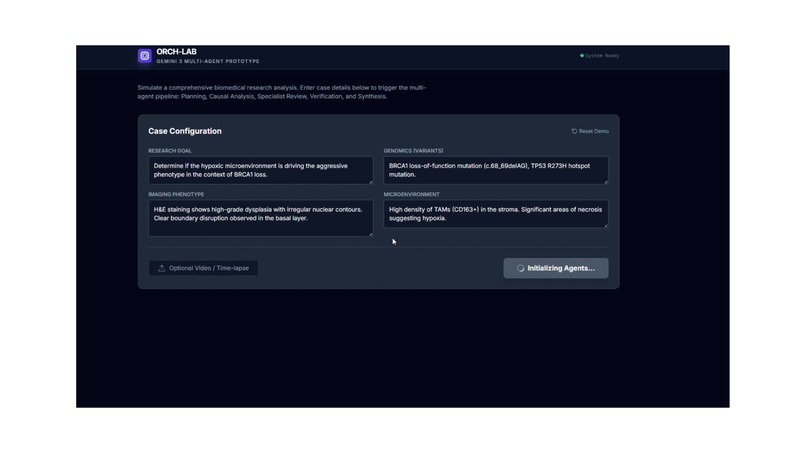
Ideated how Gemini 3 could accelerate multimodal research through orchestration and rapid prototyping.
Disclaimer & Responsible Use
Research-Only Demonstration (Not Medical Advice) This application is a research-only visualization built to demonstrate system design, orchestration, and rapid prototyping using Gemini 3 (Google AI Studio). It is NOT a medical device, and it is NOT intended for clinical use. This system does not provide: Medical advice Diagnosis Prognosis Treatment recommendations Drug dosing decisions Radiation planning Clinical decision support All outputs are non-actionable research hypotheses, generated for the purpose of technical demonstration only. No Clinical Decision-Making This work must not be used to: Guide real patient care Replace clinical judgment Replace radiologists, pathologists, or oncologists Influence medication, surgery, chemotherapy, or radiation decisions If you require medical guidance, consult a licensed healthcare professional.
Application of Gemini 3 in Biomedical Research Acceleration (Research-Only)
Inspiration
Biomedical research rarely fails because the underlying science is wrong — it fails because the system never becomes real. In oncology and translational workflows, reasoning is inherently multimodal, non-linear, and iterative, yet most AI demos still behave like thin prompt wrappers.
We wanted to explore Gemini 3 as something more useful: not a chatbot, but a research orchestrator that can:
plan → route → execute → validate → detect contradictions → synthesize
…while remaining strictly research-only.
What it does
Application of Gemini 3 in Biomedical Research Acceleration is a research-only system visualization demonstrating how Gemini 3 AI Studio can accelerate biomedical system design across three case studies:
- Orchestration — multi-agent research workflows with planning + routing
- Rapid prototyping — compressing the “Idea → Agent System → Iteration” loop
- Multimodal orchestration — spatial-temporal reasoning across modalities (imaging, genomics, PK/PD, toxicity, literature)
In addition, we ideated and built 10+ biomedical interface prototypes, spanning abstract-to-applied concepts — from a virtual digital cell twin to an agentic virtual tumor board — all framed as research infrastructure rather than clinical tools.
How we built it
Core design principle: Structured research artifacts (not chat)
A key decision was to treat every intermediate output as a machine-usable artifact, not free-form prose.
Each specialist agent produces strict JSON constrained by Pydantic schemas, including:
- structured fields
- confidence values
- limitations and uncertainty
- research-only framing
This makes the system behave like research infrastructure rather than a prompt chain.
Core architecture: DAG Orchestrator (Gemini 3)
1) Planning + routing (dynamic workflow execution)
Gemini 3 generates a Directed Acyclic Graph (DAG) execution plan per query. The plan determines:
- which specialist agents to run
- dependency ordering
- what can run in parallel
- which modalities are present or missing
This avoids brittle fixed pipelines and enables dynamic routing.
2) Specialist agents (modular, scoped roles)
We ideated and worked upon modular specialist agents, each responsible for a single modality or research function:
- Literature Evidence Pack Agent (RAG-style)
- Genomics Interpretation Agent
- Imaging Phenotype Agent
- PK/PD Reasoning Agent (synthetic curve demo)
- Toxicity + Survival Curve Agent (synthetic curve demo)
- Research Intervention Categorization Agent (explicitly non-clinical)
Each agent is intentionally narrow, with schema constraints to prevent scope creep and reduce hallucination risk.
3) Validation + governance layer
After each agent runs, Gemini 3 executes a validator stage enforcing:
- schema correctness
- research-only compliance
- hallucination risk checks
- uncertainty + limitations reporting
- reduction of overconfident or clinical language
This prevents weak intermediate outputs from silently contaminating the final synthesis.
4) Conflict detection + synthesis
Once all validated outputs exist, Gemini 3 runs:
- cross-agent conflict detection (contradictions surfaced explicitly)
final synthesis, producing a structured research artifact containing:
- integrated hypothesis map
- evidence alignment across modalities
- uncertainty + missing data
- surfaced contradictions
- proposed validation experiments
This is what makes the system orchestration rather than prompt chaining.
Case Studies
Case 1 — Explore Orchestration (Multi-Agent Research Workflow System)
This case demonstrates Gemini 3 as a true multi-agent orchestrator coordinating specialist roles like a research team.
What we worked upon
- A planner step where Gemini 3 generates a DAG execution plan
- Routing logic so only relevant agents run based on available modalities
- A modular set of specialist agents scoped to single domains
- A validator stage after each agent output
- A final synthesis stage producing a coherent research artifact instead of a generic answer
Why this matters Most “multi-agent” demos are just sequential prompts. This case demonstrates orchestration as an actual system backbone: workflow planning + modular execution + governance.
Case 2 — Explore Rapid Prototyping (Idea-to-Agent Loop)
This case focuses on compressing the most expensive part of biomedical innovation:
turning abstract ideas into working system prototypes
We built 10+ prototypes exploring biomedical product interfaces and orchestration patterns. The goal wasn’t to build a finished clinical product — it was to rapidly test workflows, interaction patterns, and system structure.
Prototypes explored included
Virtual digital cell twin interface An interactive concept where a researcher queries simulated cell states, explores perturbations, and generates mechanistic hypotheses (aging + cancer framing).
Agentic virtual tumor board interface A multi-specialist collaboration concept where agent roles mirror a tumor board (radiology, pathology, genomics, pharmacology), while remaining strictly research-only.
We also explored prototypes around:
Interfaces Explored (Research-Only)
To support the three case studies (system orchestration, rapid prototyping, and multimodal spatial-temporal reasoning), we explored a set of modular biomedical research interfaces aligned with our specialist agents and orchestration layers. Each interface was designed to produce structured research artifacts (not free-form chat), with explicit uncertainty and no clinical recommendations.
1) Imaging Agent Interface
An imaging interface that converts imaging-derived signals (or synthetic feature representations) into interpretable hypotheses.
2) Drug Dose Simulation Interface (PK/PD)
A PK/PD simulation interface that generates synthetic exposure curves to demonstrate how pharmacokinetic reasoning can be integrated into a larger orchestrated workflow.
3) Multimodal Research Orchestrator Interface
A multimodal orchestration interface where Gemini 3 coordinates evidence across imaging, genomics, PK/PD signals, toxicity curves, and literature evidence packs.
4) System Orchestrator Interface (DAG Planner)
A system orchestration interface that visualizes Gemini 3 generating a Directed Acyclic Graph (DAG) execution plan per research query. This interface highlights planning and routing as first-class capabilities: determining which agents to run, dependency ordering, and how missing modalities change execution—moving beyond brittle fixed pipelines.
5) Survival / Toxicity Agent Interface
A survival and toxicity visualization interface that outputs synthetic risk curves and adverse-event likelihoods for demonstration. The purpose is not prediction, but showing how an orchestrated system can represent outcome-like signals as structured artifacts and integrate them into conflict detection and synthesis.
6) EHR Context + Research Intervention Interface
An EHR-like structured context interface representing non-identifiable patient-like metadata (age bucket, tumor type, stage bucket, constraints).
7) Biomedical Literature Evidence Interface (RAG-style)
Why this matters Fast UI/UX prototyping matters in biomedical research because the biggest bottleneck is rarely the idea — it’s the time and engineering overhead required to turn the idea into a real, testable system.
In practice, a lot of promising biomedical concepts die in the gap between:
“This could work” → “We have something usable that a lab/team can iterate on.”
Unlike many software domains, biomedical workflows are messy: multimodal data, incomplete evidence, uncertainty, long iteration cycles, and high cost of validation. If you can’t prototype quickly, you can’t:
- test whether a workflow is even usable
- discover what evidence is missing
- surface contradictions early
- iterate on agent roles and system design
- translate hypotheses into validation experiments
Fast prototyping also reduces the “false confidence” problem. Instead of building a polished single-output demo, you build a system that forces real research behavior: planning, verification, uncertainty tracking, and iteration.
That’s why compressing the Idea → Agent System → Iteration loop is so valuable: it makes biomedical innovation more like building infrastructure, not just writing papers or producing static AI demos.
Case 3 — Explore Multimodal Orchestrator (Spatial-Temporal Reasoning)
This case demonstrates the most advanced capability:
multimodal reasoning across time, treated as a workflow rather than a single model response.
We worked upon a multimodal pipeline demo (Colab + Gemini 3) showing how an orchestrator can integrate:
- imaging features
- genomics variants
- PK/PD response curves
- toxicity risk curves
- literature evidence packs
- video-based spatial-temporal reasoning (frame sampling → causal graph)
Core system pattern
Gemini 3 plays multiple orchestration roles:
1) Planner Builds a DAG execution plan depending on:
- the question
- which modalities are present
- which agents/tools are allowed
2) Validator Runs after each agent output to enforce:
- schema integrity
- hypothesis-only language
- limitations + uncertainty
- no clinical recommendations
3) Conflict Detector Compares agent outputs to detect contradictions such as:
- imaging phenotype not aligning with molecular pathway hypotheses
- PK reasoning inconsistent with toxicity signals
- literature evidence conflicting with agent claims
Instead of hiding disagreement, the system surfaces it as a first-class output.
4) Final Synthesizer Produces a structured research artifact containing:
- integrated hypothesis map
- evidence links across modalities
- conflicts + resolution experiments
- uncertainty propagation
- missing evidence checklist
Why this matters Most healthcare multimodal demos treat multimodal as “text + image input.” This case treats multimodal as process reasoning: timelines, transitions, uncertainty, and evidence consistency.
Research-only compliance built into the system
Because this project touches biomedical workflows, we treated safety as architecture — not a disclaimer.
Across all three cases:
- outputs are framed as non-actionable research hypotheses
- schemas enforce limitations + uncertainty fields
- validator steps reduce clinical language
- the system refuses clinical decision-making
Challenges we ran into
- Designing a system that supports backward-loop thinking (revise, re-run, request missing evidence) rather than linear input → output
- Preventing “false coherence” when agents disagree across modalities
- Making multimodal reasoning handle time + uncertainty, not just multiple input types
- Enforcing research-only boundaries as a system property, not just a disclaimer
- Keeping 10+ prototypes aligned under one coherent system narrative
Accomplishments
- Worked upon a prototyped orchestration abstraction using DAG planning + routing, not static agent sequencing
- Worked upon a research-grade conceptual workflow: validate → conflict detect → synthesize
- Developed 10+ biomedical prototypes showing how abstract ideas can become working system designs quickly
- Delivered a visualization that communicates architecture clearly (agents, dependencies, validation, synthesis)
What we learned
- Multi-agent systems fail without orchestration: planning, routing, validation, and conflict detection are the difference between a demo and a system
- In biomedical workflows, the most valuable capability is not “answering” — it’s evidence alignment across modalities
- Verification is non-optional: the weakest agent can poison the final synthesis unless outputs are gated
- Multimodal reasoning is primarily about process, time, transitions, and uncertainty propagation, not just image understanding
Closing note
This project is intentionally research-only. It is a system visualization showing how Gemini 3 can act as orchestrated research infrastructure — generating structured intermediate artifacts, surfacing contradictions, and proposing validation experiments — without producing clinical recommendations.
Short Summary
Pipeline 1 uses Gemini 3 as a system orchestrator to plan and run a DAG of specialist agents (RAG, genomics, imaging, PK/PD, toxicity), validate their outputs, detect conflicts, and synthesize a final research report, while Pipeline 2 uses Gemini 3 as a multimodal research orchestrator to reason directly over raw multimodal inputs (especially video) by extracting a time-ordered event timeline and causal graph, then verifying and synthesizing cross-modal hypotheses.
Built With
- agenticai
- aiagent
- colab
- figma
- gemini3
- googleaistudio
- python




Log in or sign up for Devpost to join the conversation.